Descriptive statistics
Last compiled on oktober, 2022
We will use the RSiena objects of clubs to describe the kudos networks, constant covariates and behavioral variables.
Getting started
Clean the working environment and load in the club data.
# clean the working environment
rm (list = ls( ))
# load the RSiena objects
load("clubdata_rsiena_freq.RData")
load("clubdata_rsiena_vol.RData")
# and the list containing club data-setes
load("clubdata.RData")general custom functions
fpackage.check: Check if packages are installed (and install if not) in R (source)fload.R: function to load R-objects under new names.
fpackage.check <- function(packages) {
lapply(packages, FUN = function(x) {
if (!require(x, character.only = TRUE)) {
install.packages(x, dependencies = TRUE)
library(x, character.only = TRUE)
}
})
}
fload.R <- function(fileName){
load(fileName)
get(ls()[ls() != "fileName"])
}necessary packages
We install and load the packages we need later on:
RSiena: RSiena models, some descriptives on network leveligraph: Descriptives (dyad/triad census, degrees)tidyversetidyr: for tidy datamoments: for calculating statistics (e.g., kurtosis, standard error)dplyr: for data manipulationggplot2: for data visualizationforcats: for handling categorical variablknitr: for generating tableskableExtra: for manipulating tablesnetwork: for network analysissna: for network analysis
packages = c("RSiena", "igraph", "tidyverse", "tidyr", "moments", "dplyr", "ggplot2", "forcats", "knitr", "kableExtra", "network", "sna")
fpackage.check(packages)additional functions
Now define some additional functions we use later on to describe our data (see www.jochemtolsma.nl).
fdensity: calculate density (exclude NA and structural zeros)
fdensityintra: calculate density within group (exclude NA and structural zeros)
fdensityinter: calculate density between groups (exclude NA and structural zeros)
fhomomat: based on ego/alter characteristics, construct dyad characteristic whether or not ego/alter are samefndyads: calculate all valid dyads (no NA or structural zeros)
fscolnet: calculate Coleman’s segregation index on the network-level
fMoran.i: calculate Moran’s I spatial autocorrelation statistic (see here)
# density: observed relations divided by possible relations
fdensity <- function(x) {
# x is your nomination network make sure diagonal cells are NA
diag(x) <- NA
# take care of RSiena structural zeros, set as missing.
x[x == 10] <- NA
sum(x == 1, na.rm = T)/(sum(x == 1 | x == 0, na.rm = T))
}
# calculate intragroup density
fdensityintra <- function(x, A) {
# A is matrix indicating whether nodes in dyad have same node attributes
diag(x) <- NA
x[x == 10] <- NA
diag(A) <- NA
sum(x == 1 & A == 1, na.rm = T)/(sum((x == 1 | x == 0) & A == 1, na.rm = T))
}
# calculate intragroup density
fdensityinter <- function(x, A) {
# A is matrix indicating whether nodes in dyad have same node attributes
diag(x) <- NA
x[x == 10] <- NA
diag(A) <- NA
sum(x == 1 & A != 1, na.rm = T)/(sum((x == 1 | x == 0) & A != 1, na.rm = T))
}
# construct dyad characteristic whether nodes are similar/homogenous
fhomomat <- function(x) {
# x is a vector of node-covariate
xmat <- matrix(x, nrow = length(x), ncol = length(x))
xmatt <- t(xmat)
xhomo <- xmat == xmatt
return(xhomo)
}
# a function to calculate all valid dyads.
fndyads <- function(x) {
diag(x) <- NA
x[x == 10] <- NA
(sum((x == 1 | x == 0), na.rm = T))
}
# a function to calculate all valid intragroupdyads.
fndyads2 <- function(x, A) {
diag(x) <- NA
x[x == 10] <- NA
diag(A) <- NA
(sum((x == 1 | x == 0) & A == 1, na.rm = T))
}
fscolnet <- function(network, ccovar) {
# Calculate coleman on network level:
# https://reader.elsevier.com/reader/sd/pii/S0378873314000239?token=A42F99FF6E2B750436DD2CB0DB7B1F41BDEC16052A45683C02644DAF88215A3379636B2AA197B65941D6373E9E2EE413
fhomomat <- function(x) {
xmat <- matrix(x, nrow = length(x), ncol = length(x))
xmatt <- t(xmat)
xhomo <- xmat == xmatt
return(xhomo)
}
fsumintra <- function(x, A) {
# A is matrix indicating whether nodes constituting dyad have same characteristics
diag(x) <- NA
x[x == 10] <- NA
diag(A) <- NA
sum(x == 1 & A == 1, na.rm = T)
}
# expecation w*=sum_g sum_i (ni((ng-1)/(N-1)))
network[network == 10] <- NA
ni <- rowSums(network, na.rm = T)
ng <- NA
for (i in 1:length(ccovar)) {
ng[i] <- table(ccovar)[rownames(table(ccovar)) == ccovar[i]]
}
N <- length(ccovar)
wexp <- sum(ni * ((ng - 1)/(N - 1)), na.rm = T)
# wgg1 how many intragroup ties
w <- fsumintra(network, fhomomat(ccovar))
Scol_net <- ifelse(w >= wexp, (w - wexp)/(sum(ni, na.rm = T) - wexp), (w - wexp)/wexp)
return(Scol_net)
}
fMoran.I <- function(x, weight, scaled = FALSE, na.rm = FALSE, alternative = "two.sided", rowstandardize = TRUE) {
if (rowstandardize) {
if (dim(weight)[1] != dim(weight)[2])
stop("'weight' must be a square matrix")
n <- length(x)
if (dim(weight)[1] != n)
stop("'weight' must have as many rows as observations in 'x'")
ei <- -1/(n - 1)
nas <- is.na(x)
if (any(nas)) {
if (na.rm) {
x <- x[!nas]
n <- length(x)
weight <- weight[!nas, !nas]
} else {
warning("'x' has missing values: maybe you wanted to set na.rm = TRUE?")
return(list(observed = NA, expected = ei, sd = NA, p.value = NA))
}
}
ROWSUM <- rowSums(weight)
ROWSUM[ROWSUM == 0] <- 1
weight <- weight/ROWSUM
s <- sum(weight)
m <- mean(x)
y <- x - m
cv <- sum(weight * y %o% y)
v <- sum(y^2)
obs <- (n/s) * (cv/v)
if (scaled) {
i.max <- (n/s) * (sd(rowSums(weight) * y)/sqrt(v/(n - 1)))
obs <- obs/i.max
}
S1 <- 0.5 * sum((weight + t(weight))^2)
S2 <- sum((apply(weight, 1, sum) + apply(weight, 2, sum))^2)
s.sq <- s^2
k <- (sum(y^4)/n)/(v/n)^2
sdi <- sqrt((n * ((n^2 - 3 * n + 3) * S1 - n * S2 + 3 * s.sq) - k * (n * (n - 1) * S1 - 2 * n *
S2 + 6 * s.sq))/((n - 1) * (n - 2) * (n - 3) * s.sq) - 1/((n - 1)^2))
alternative <- match.arg(alternative, c("two.sided", "less", "greater"))
pv <- pnorm(obs, mean = ei, sd = sdi)
if (alternative == "two.sided")
pv <- if (obs <= ei)
2 * pv else 2 * (1 - pv)
if (alternative == "greater")
pv <- 1 - pv
list(observed = obs, expected = ei, sd = sdi, p.value = pv)
} else {
if (dim(weight)[1] != dim(weight)[2])
stop("'weight' must be a square matrix")
n <- length(x)
if (dim(weight)[1] != n)
stop("'weight' must have as many rows as observations in 'x'")
ei <- -1/(n - 1)
nas <- is.na(x)
if (any(nas)) {
if (na.rm) {
x <- x[!nas]
n <- length(x)
weight <- weight[!nas, !nas]
} else {
warning("'x' has missing values: maybe you wanted to set na.rm = TRUE?")
return(list(observed = NA, expected = ei, sd = NA, p.value = NA))
}
}
# ROWSUM <- rowSums(weight) ROWSUM[ROWSUM == 0] <- 1 weight <- weight/ROWSUM
s <- sum(weight)
m <- mean(x)
y <- x - m
cv <- sum(weight * y %o% y)
v <- sum(y^2)
obs <- (n/s) * (cv/v)
if (scaled) {
i.max <- (n/s) * (sd(rowSums(weight) * y)/sqrt(v/(n - 1)))
obs <- obs/i.max
}
S1 <- 0.5 * sum((weight + t(weight))^2)
S2 <- sum((apply(weight, 1, sum) + apply(weight, 2, sum))^2)
s.sq <- s^2
k <- (sum(y^4)/n)/(v/n)^2
sdi <- sqrt((n * ((n^2 - 3 * n + 3) * S1 - n * S2 + 3 * s.sq) - k * (n * (n - 1) * S1 - 2 * n *
S2 + 6 * s.sq))/((n - 1) * (n - 2) * (n - 3) * s.sq) - 1/((n - 1)^2))
alternative <- match.arg(alternative, c("two.sided", "less", "greater"))
pv <- pnorm(obs, mean = ei, sd = sdi)
if (alternative == "two.sided")
pv <- if (obs <= ei)
2 * pv else 2 * (1 - pv)
if (alternative == "greater")
pv <- 1 - pv
list(observed = obs, expected = ei, sd = sdi, p.value = pv)
}
}We cover the following:
- club characteristics
- network structure in Kudos networks
- gender composition / segregation
- behavior: activity level (frequency and duration) and correlation
- spatial network autocorrelation: behavioral similarity in networks
Print report
Make sure to check the output of the ‘print01Report()’ function for general data descripton (degrees, network size, etc.) and a general overview of the dataset. Output is printed in a .txt file in the directory specified.
Club 1
df <- clubdata_rsiena_freq[[1]] # grab club
print01Report(df, modelname="files/club1")Club 2
df <- clubdata_rsiena_freq[[2]] # grab club
print01Report(df, modelname="files/club2")Club 3
df <- clubdata_rsiena_freq[[3]] # grab club
print01Report(df, modelname="files/club3")Club 4
df <- clubdata_rsiena_freq[[4]] # grab club
print01Report(df, modelname="files/club4")Club 5
df <- clubdata_rsiena_freq[[5]] # grab club
print01Report(df, modelname="files/club5")Club characteristics
Some club characteristics. We show the size of the network (the number of actors) for each club and the number of active members currently (18-1-2021) on Strava, by adding to www.strava.com/clubs/… the original club id.
# attrieve from the clubdata the number of actors in each network
netsize <- c(
length(clubdata_rsiena_freq[[1]]$nodeSets$Actors),
length(clubdata_rsiena_freq[[2]]$nodeSets$Actors),
length(clubdata_rsiena_freq[[3]]$nodeSets$Actors),
length(clubdata_rsiena_freq[[4]]$nodeSets$Actors),
length(clubdata_rsiena_freq[[5]]$nodeSets$Actors))
clubsize <- c(66, 127, 373, 15, 169) # find the number of members currently on Strava
df <- data.frame( netsize = netsize, clubsize = clubsize )
print(df)#> netsize clubsize
#> 1 27 66
#> 2 58 127
#> 3 159 373
#> 4 9 15
#> 5 76 169Kudos network
Now let’s describe the Kudos network.
1. Node-level
Starting, again, with indegrees and outdegrees: who receives and gives Kudos? We take from the RSiena object the Kudos network for each club, subset the first wave, and turn it into an igraph object.
Club 1
df <- clubdata_rsiena_freq[[1]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# find in- and outdegree for each node
hist(igraph::degree(G1, mode="out"), xlab="outdegree", main="histogram of Kudo outdegree")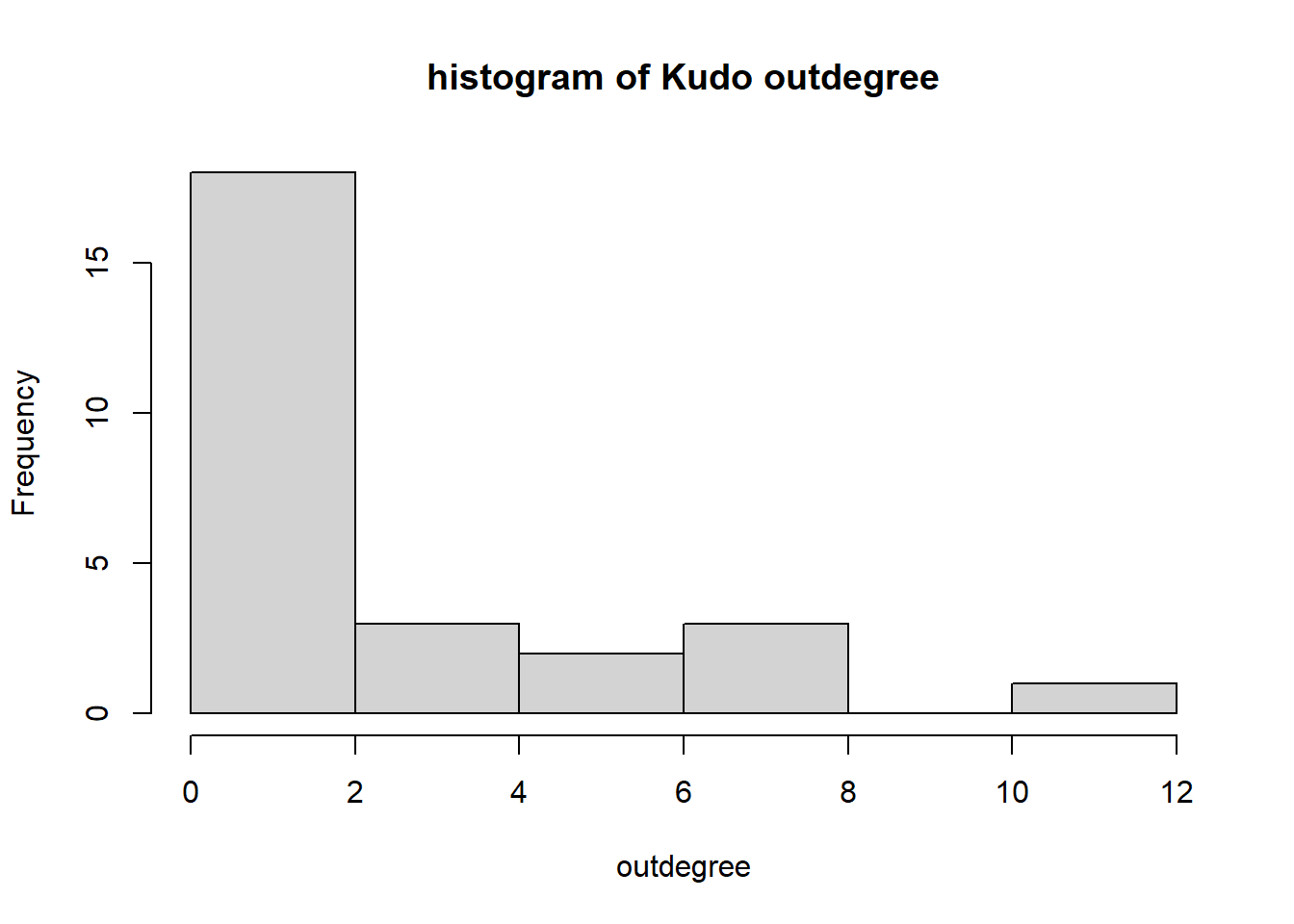
hist(igraph::degree(G1, mode="in"), xlab="indegree", main="histogram of Kudo indegree")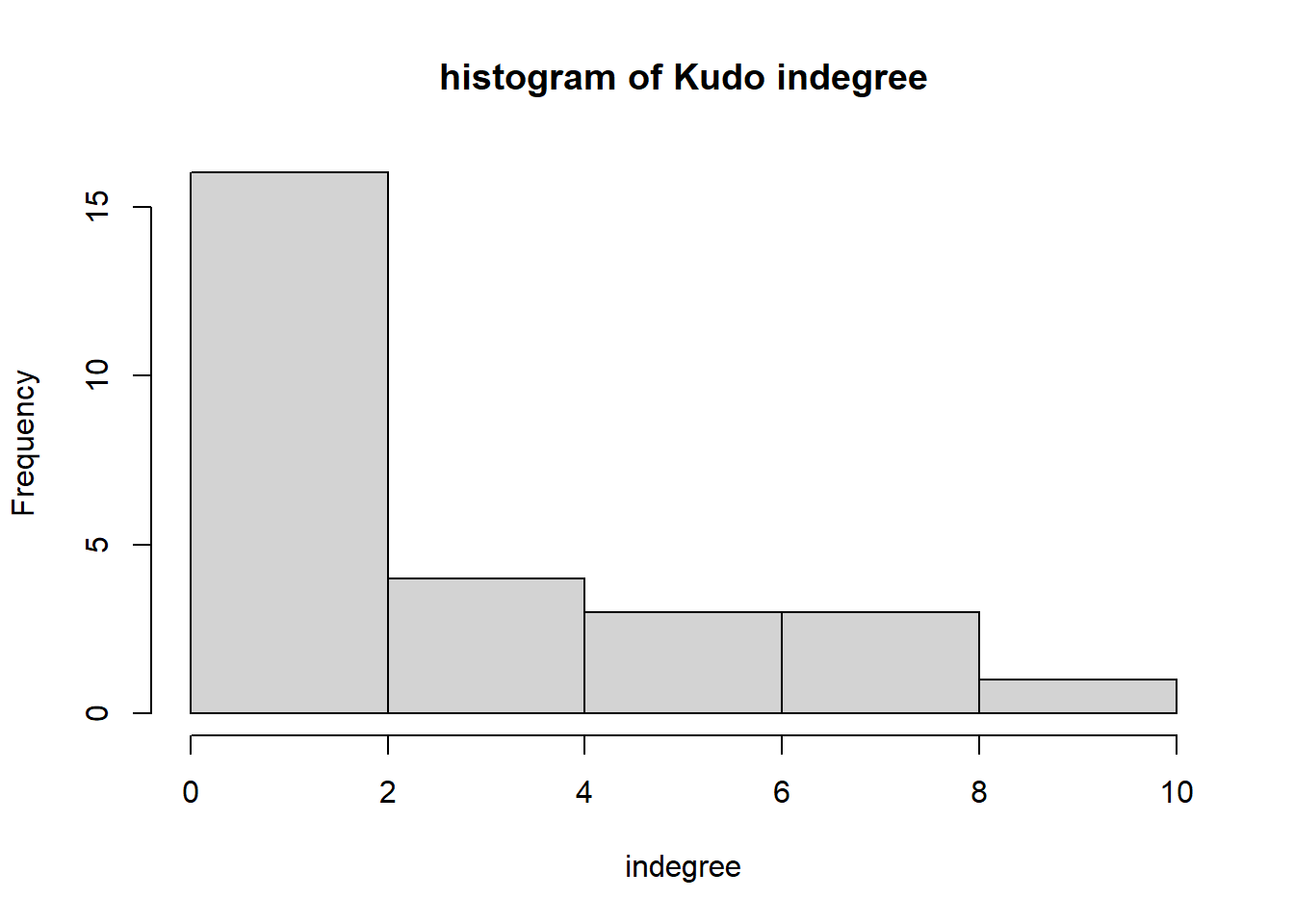
Club 2
df <- clubdata_rsiena_freq[[2]] # grab club t
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# find in- and outdegree for each node
hist(igraph::degree(G1, mode="out"), xlab="outdegree", main="histogram of Kudo outdegree")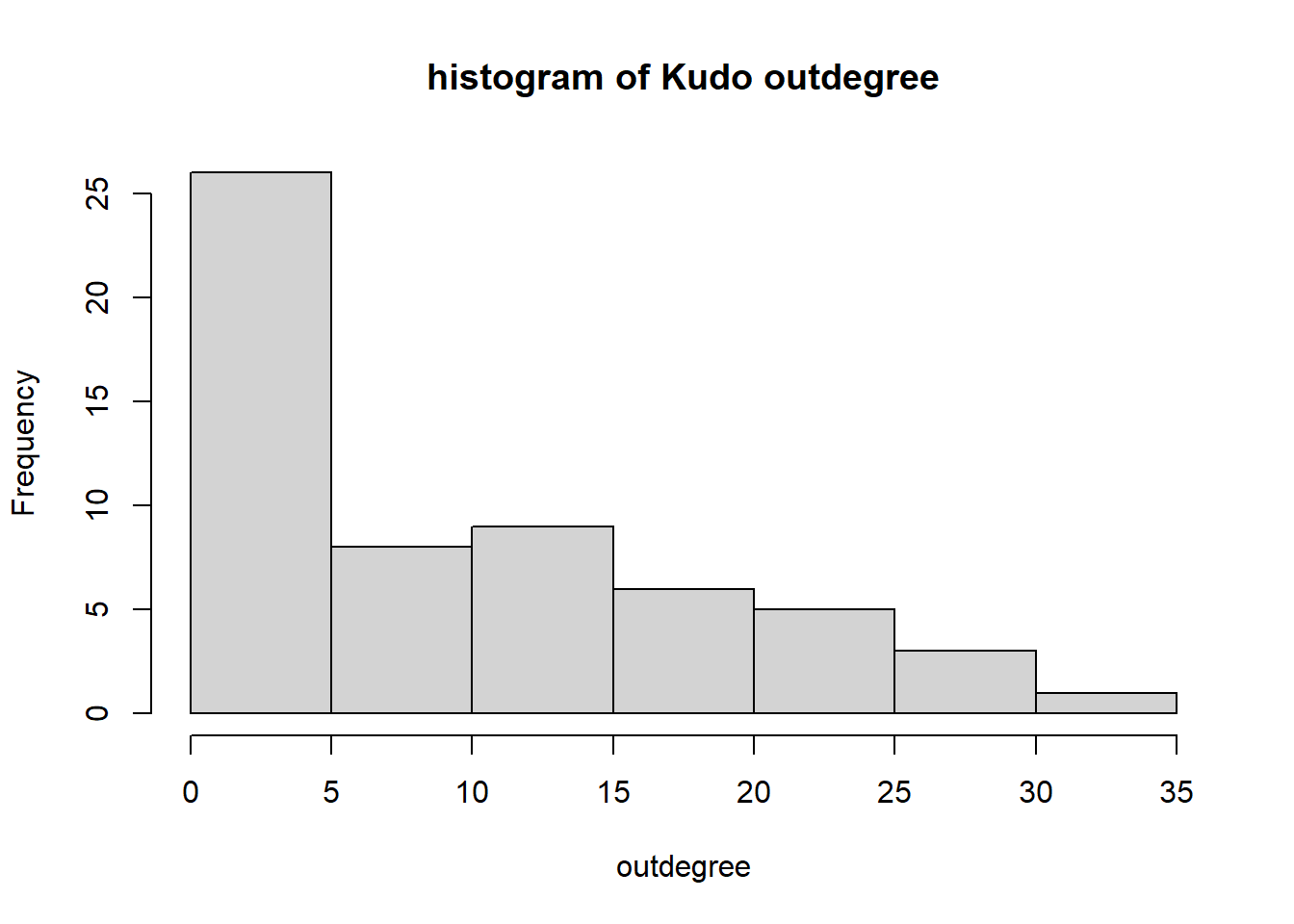
hist(igraph::degree(G1, mode="in"), xlab="indegree", main="histogram of Kudo indegree")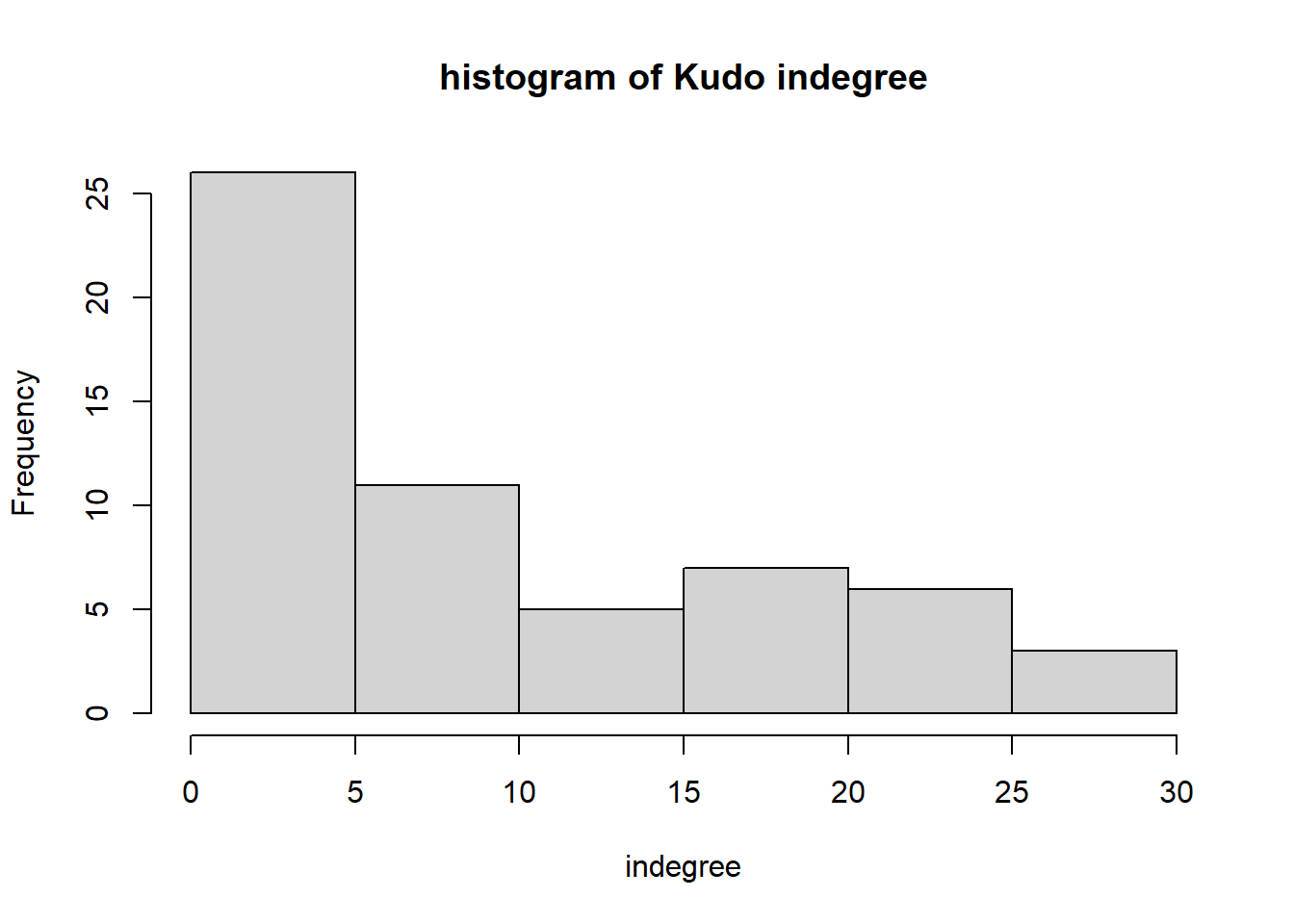
Club 3
df <- clubdata_rsiena_freq[[3]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# find in- and outdegree for each node
hist(igraph::degree(G1, mode="out"), xlab="outdegree", main="histogram of Kudo outdegree")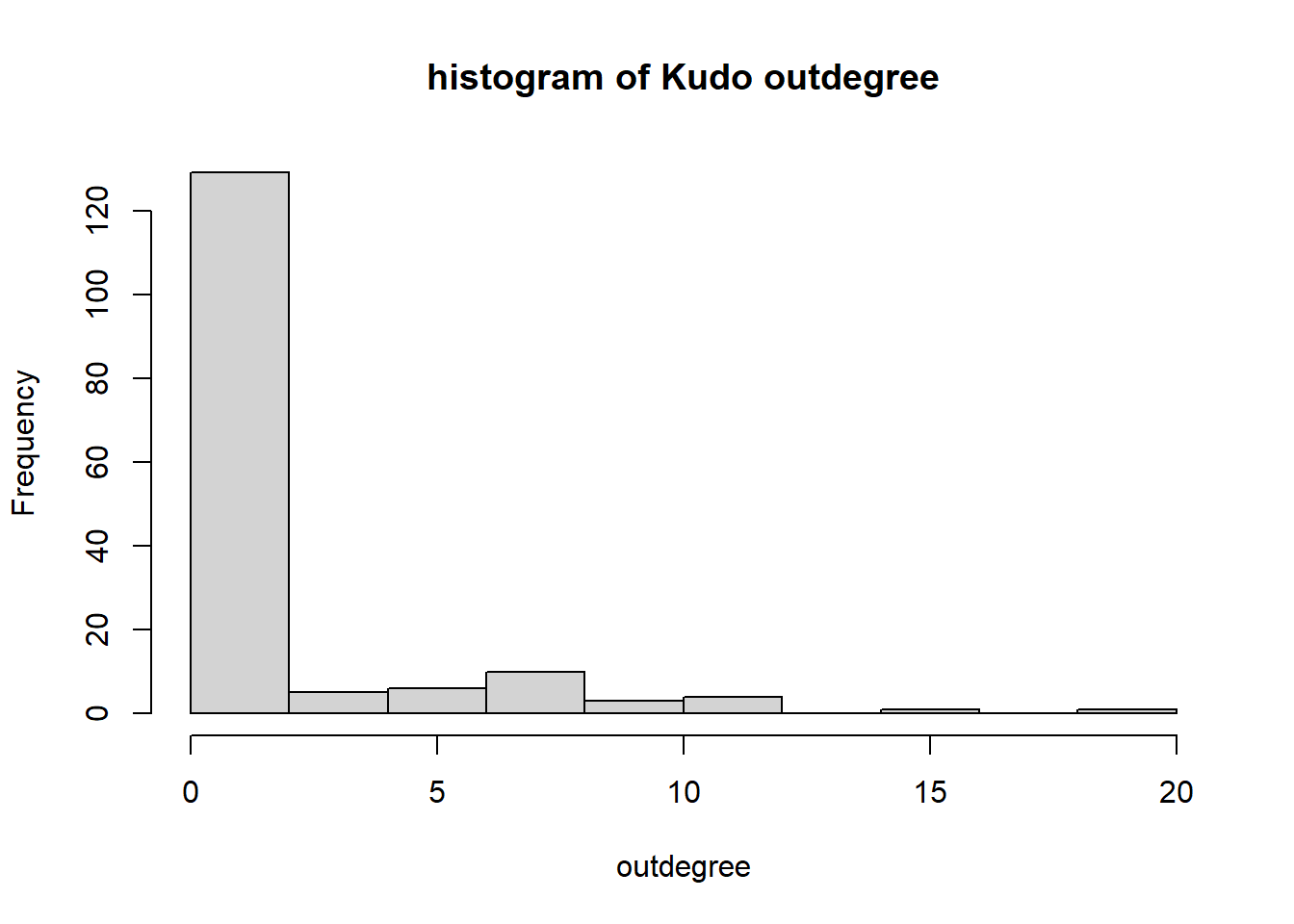
hist(igraph::degree(G1, mode="in"), xlab="indegree", main="histogram of Kudo indegree")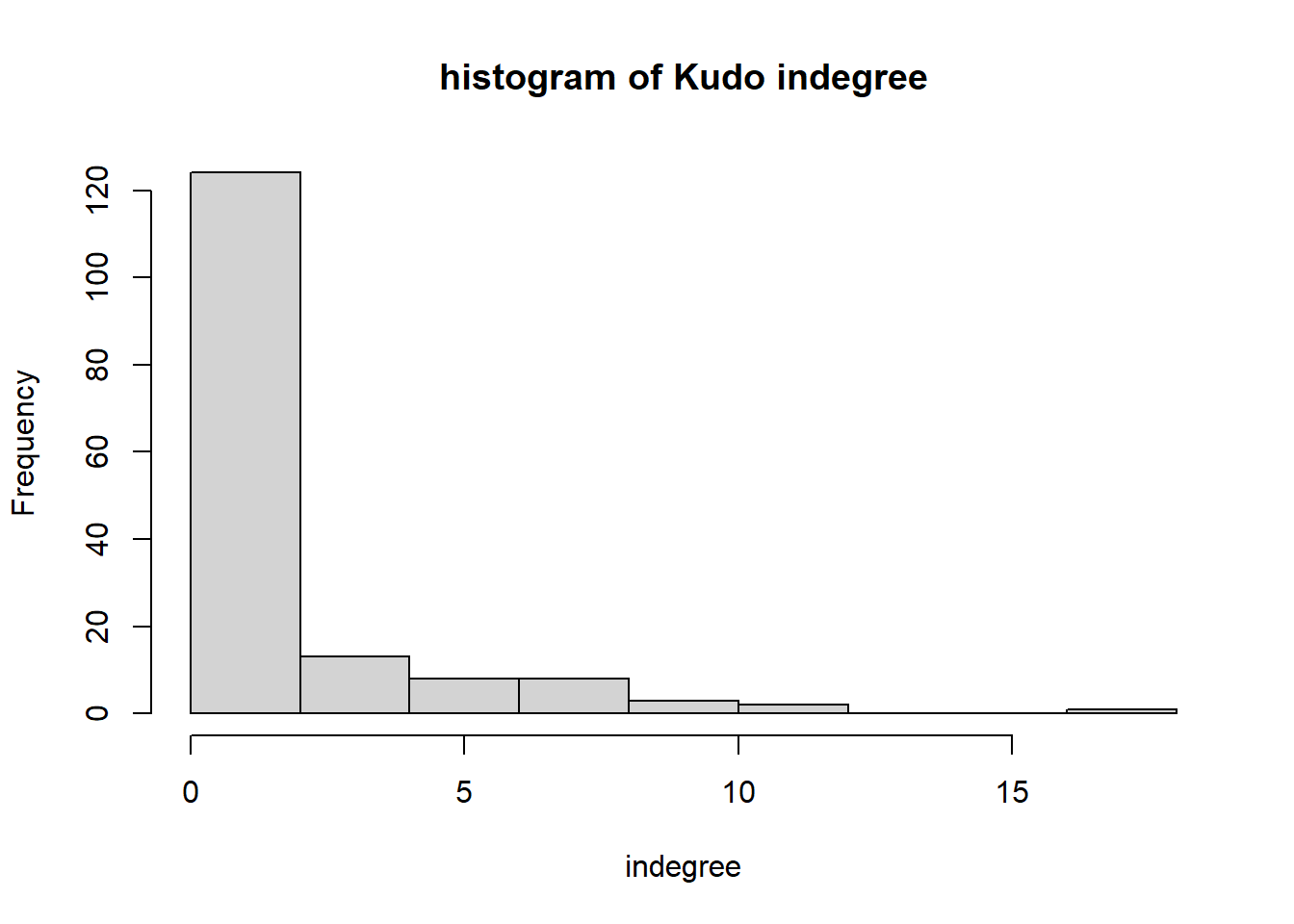
Club 4
df <- clubdata_rsiena_freq[[4]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# find in- and outdegree for each node
hist(igraph::degree(G1, mode="out"), xlab="outdegree", main="histogram of Kudo outdegree")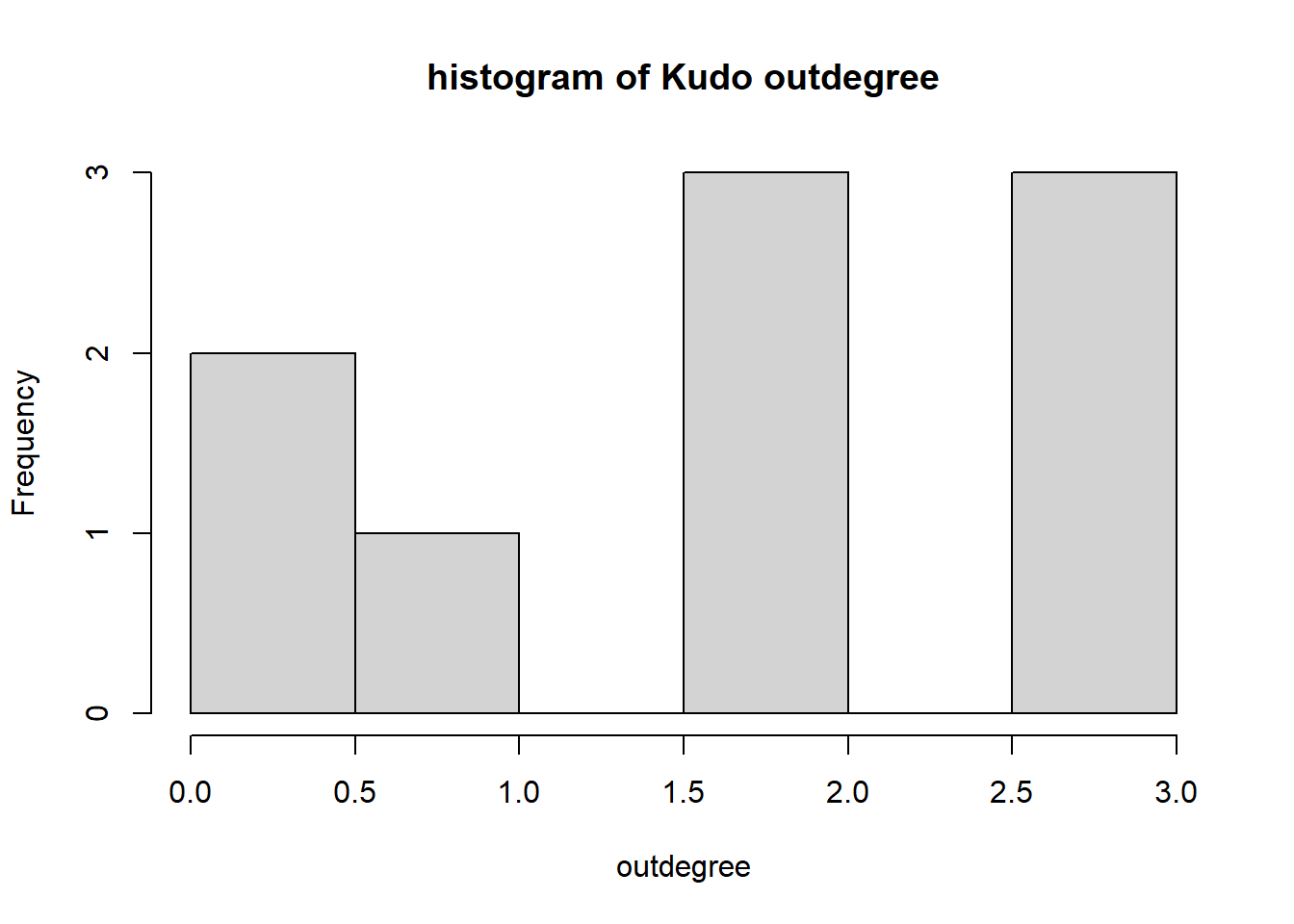
hist(igraph::degree(G1, mode="in"), xlab="indegree", main="histogram of Kudo indegree")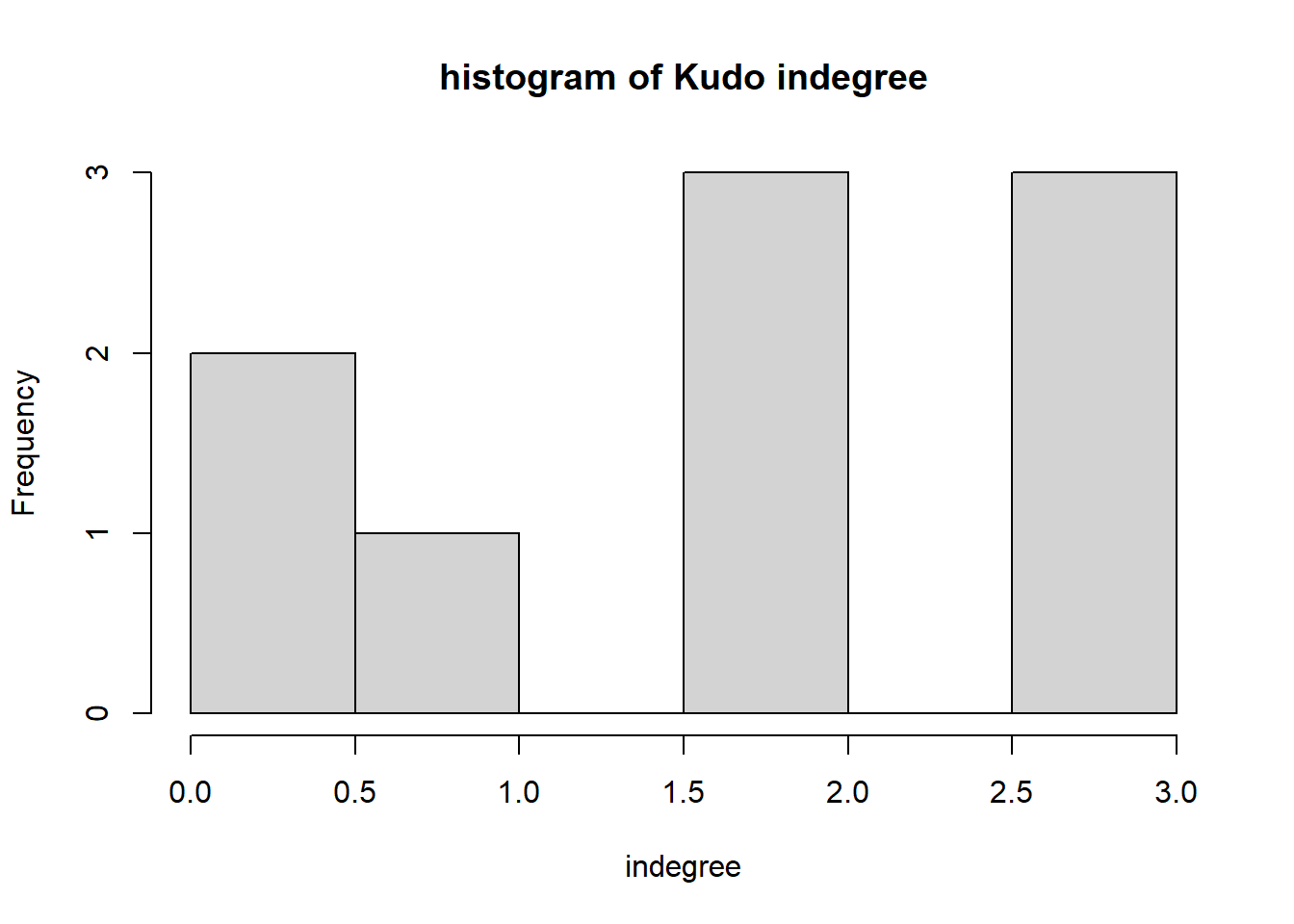
Club 5
df <- clubdata_rsiena_freq[[5]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# find in- and outdegree for each node
hist(igraph::degree(G1, mode="out"), xlab="outdegree", main="histogram of Kudo outdegree")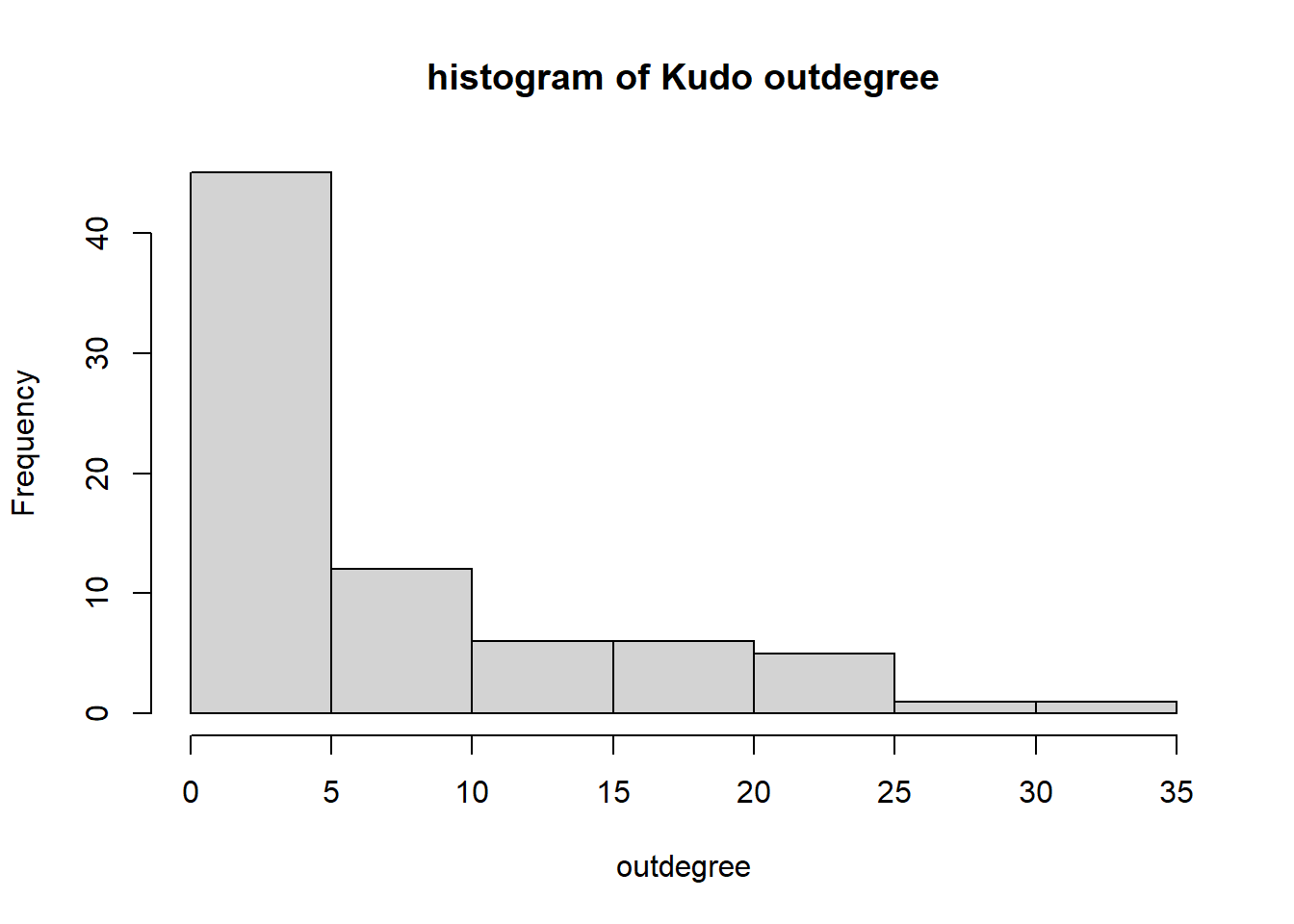
hist(igraph::degree(G1, mode="in"), xlab="indegree", main="histogram of Kudo indegree")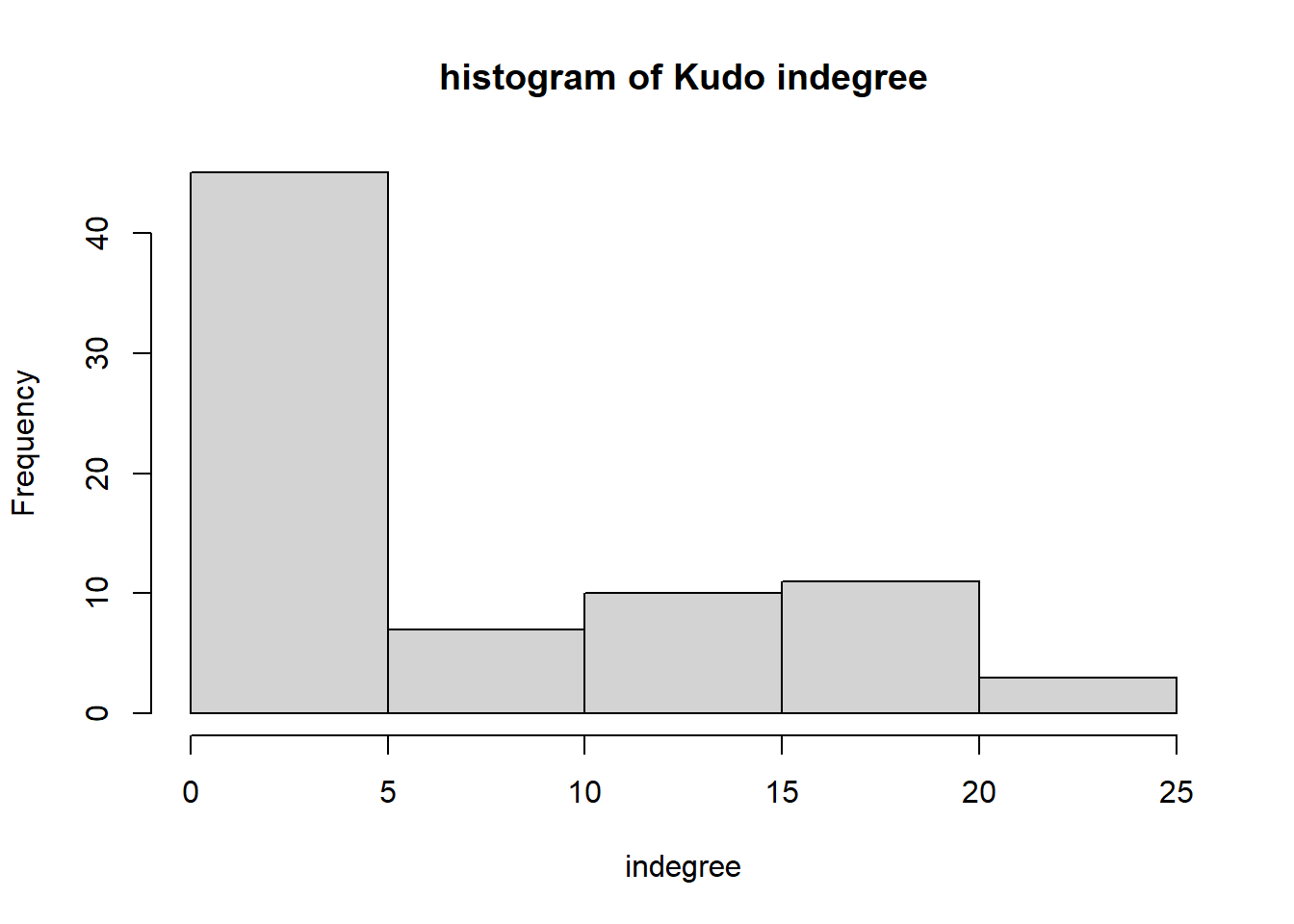
We can observe a Pareto-like-pattern: some give/receive most of the Kudos given, while most give/receive few.
2. Dyad-level
At the dyad-level: let’s see to what extent Kudos tend to be reciprocated between actors.
Club 1
# make igraph object for the club, at wave 1
df <- clubdata_rsiena_freq[[1]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# classify dyads
dyadcount <- igraph::dyad.census(G1)
# add the total number of dyads to the graph
dyadcount$total <- (vcount(G1)*(vcount(G1)-1))/2
dyadcount#> $mut
#> [1] 30
#>
#> $asym
#> [1] 5
#>
#> $null
#> [1] 316
#>
#> $total
#> [1] 351# compare values with a random graph of the same size with the same density
dens <- igraph::graph.density(G1)
size <- igraph::vcount(G1)
trial <- 1000
recip <- rep(NA, trial)
for ( i in 1:trial ){
random_graph <- igraph::erdos.renyi.game(n = size, p.or.m = dens, directed = TRUE)
recip[i] <- igraph::dyad.census(random_graph)$mut
}
{hist(recip, main="number of reciprocated Kudos in random graph", xlab="", )
abline(v=dyadcount$mut, col="red", lwd=3)}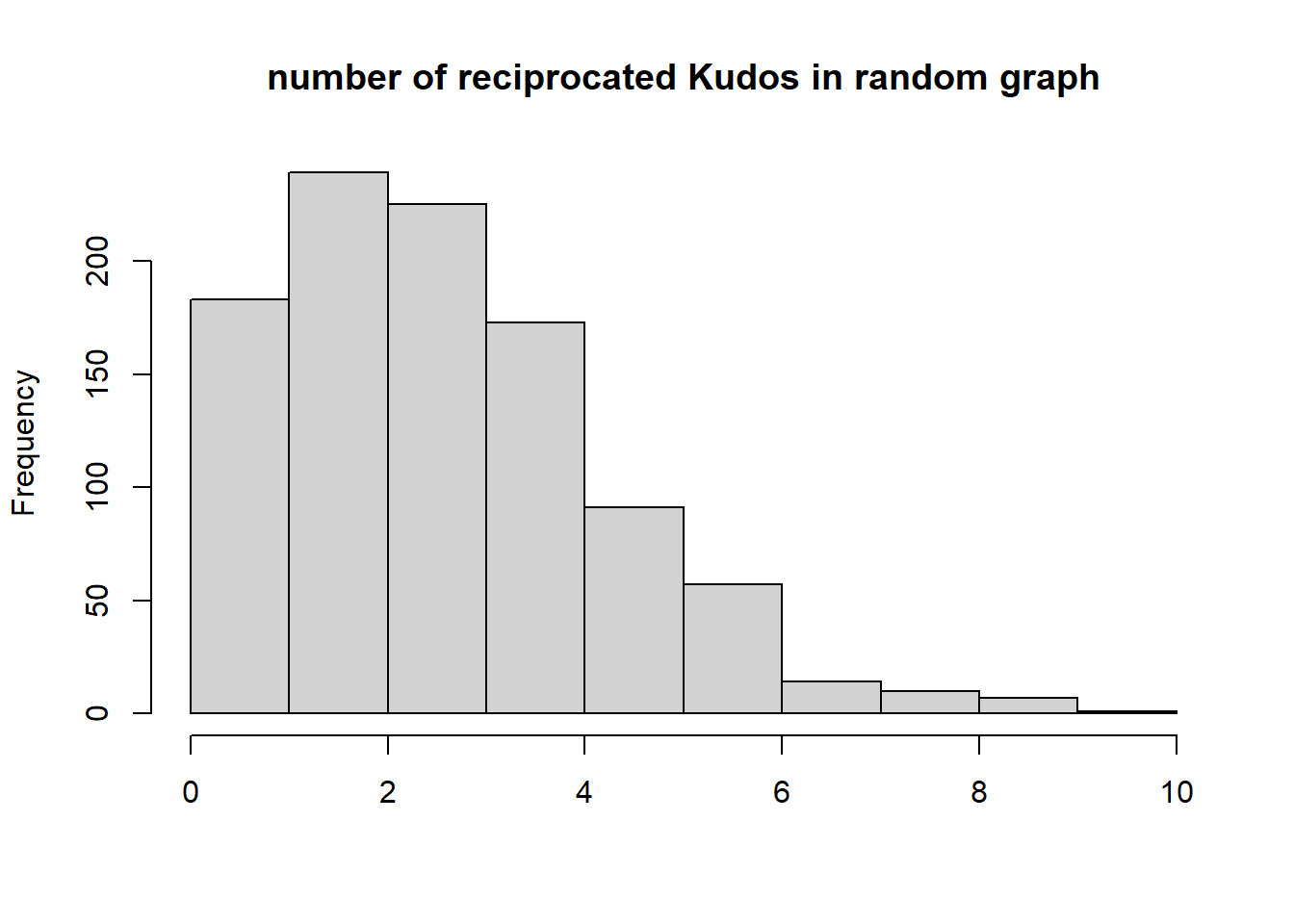
Club 2
# make igraph object for the club, at wave 1
df <- clubdata_rsiena_freq[[2]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# classify dyads
dyadcount <- igraph::dyad.census(G1)
# add the total number of dyads to the graph
dyadcount$total <- (vcount(G1)*(vcount(G1)-1))/2
dyadcount#> $mut
#> [1] 231
#>
#> $asym
#> [1] 99
#>
#> $null
#> [1] 1323
#>
#> $total
#> [1] 1653# compare values with a random graph of the same size with the same density
dens <- igraph::graph.density(G1)
size <- igraph::vcount(G1)
trial <- 1000
recip <- rep(NA, trial)
for ( i in 1:trial ){
random_graph <- igraph::erdos.renyi.game(n = size, p.or.m = dens, directed = TRUE)
recip[i] <- igraph::dyad.census(random_graph)$mut
}
{hist(recip, main="number of reciprocated Kudos in random graph", xlab="", )
abline(v=dyadcount$mut, col="red", lwd=3)}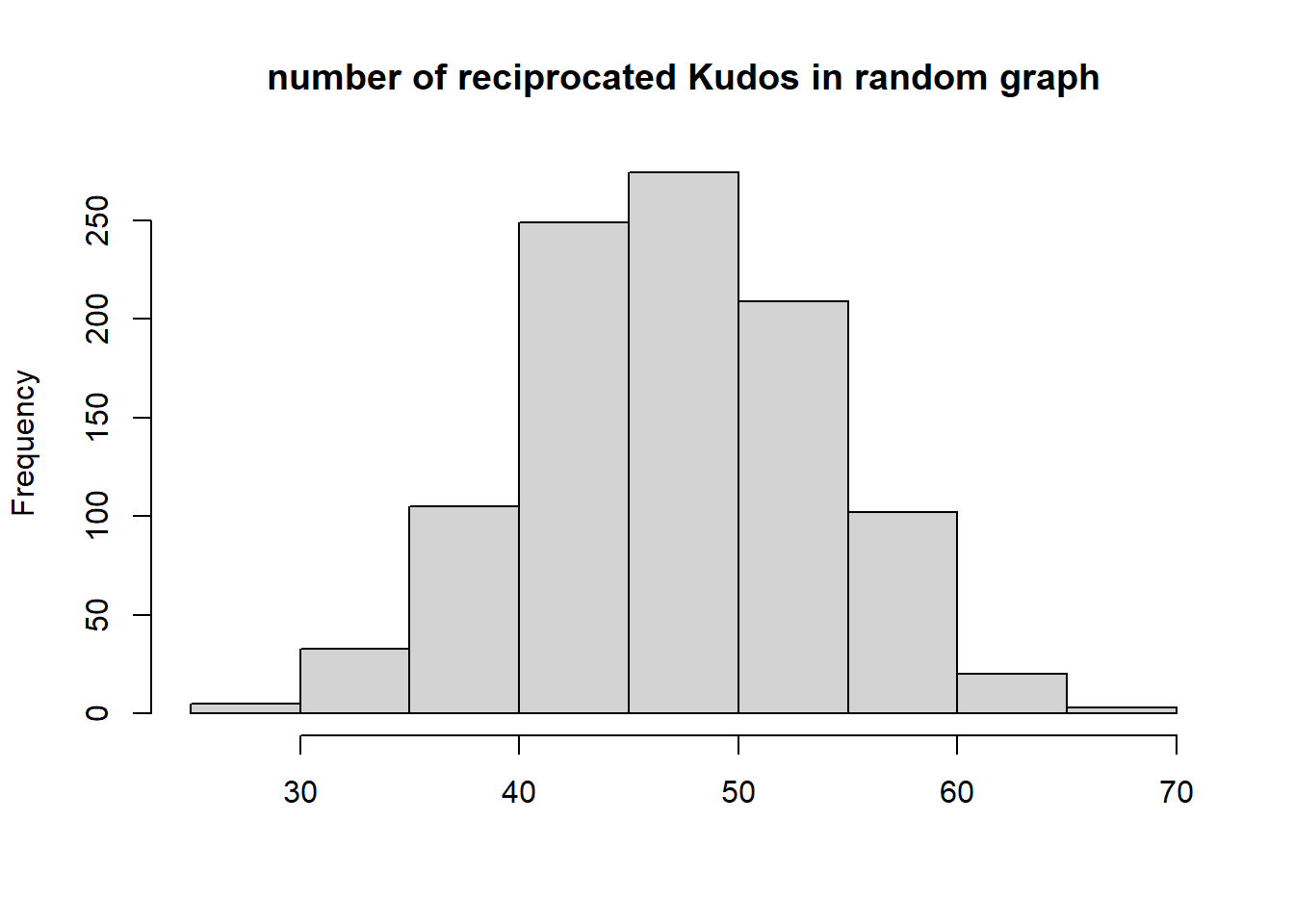
Club 3
# make igraph object for the club, at wave 1
df <- clubdata_rsiena_freq[[3]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# classify dyads
dyadcount <- igraph::dyad.census(G1)
# add the total number of dyads to the graph
dyadcount$total <- (vcount(G1)*(vcount(G1)-1))/2
dyadcount#> $mut
#> [1] 91
#>
#> $asym
#> [1] 104
#>
#> $null
#> [1] 12366
#>
#> $total
#> [1] 12561# compare values with a random graph of the same size with the same density
dens <- igraph::graph.density(G1)
size <- igraph::vcount(G1)
trial <- 1000
recip <- rep(NA, trial)
for ( i in 1:trial ){
random_graph <- igraph::erdos.renyi.game(n = size, p.or.m = dens, directed = TRUE)
recip[i] <- igraph::dyad.census(random_graph)$mut
}
{hist(recip, main="number of reciprocated Kudos in random graph", xlab="", )
abline(v=dyadcount$mut, col="red", lwd=3)}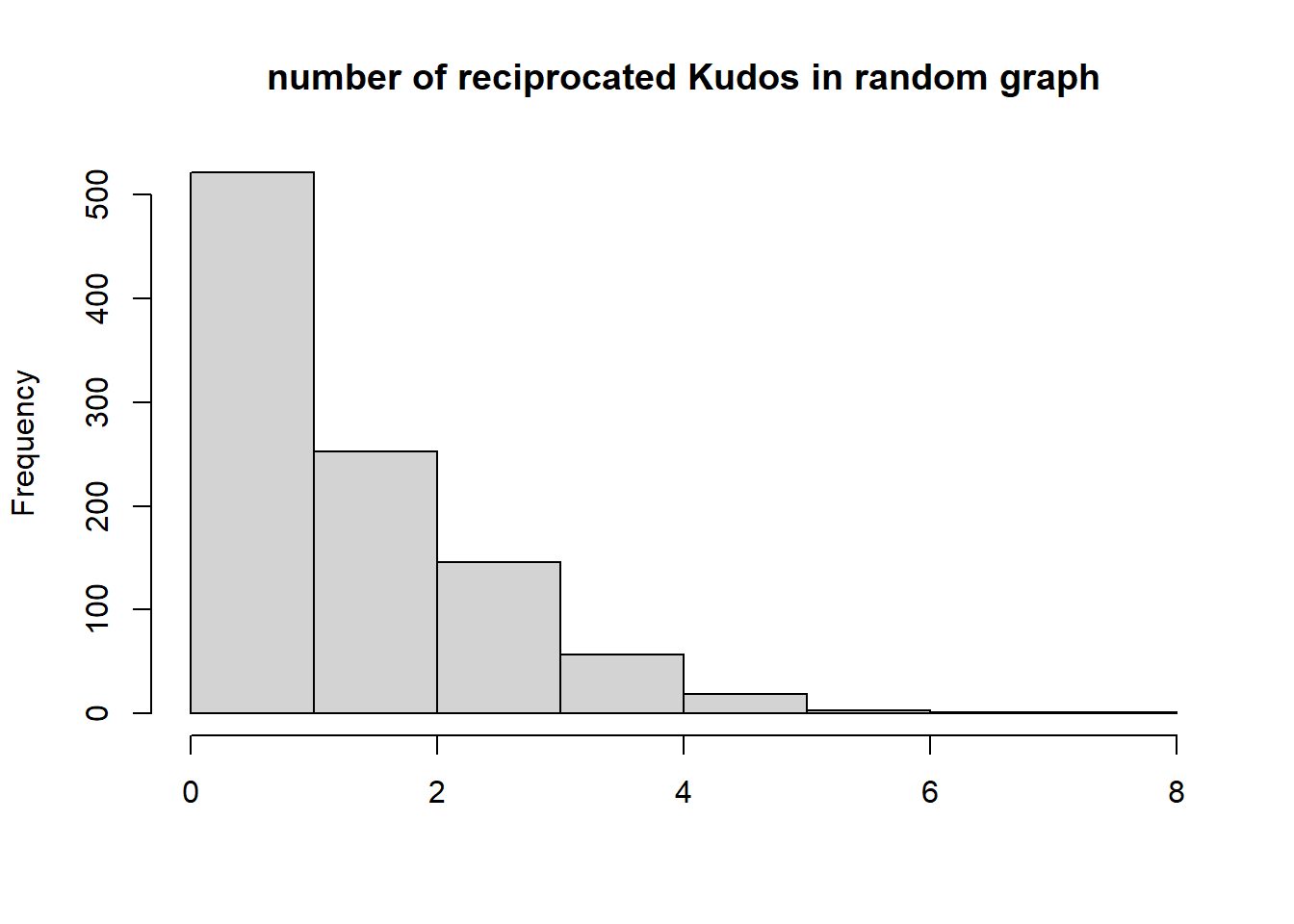
Club 4
# make igraph object for the club, at wave 1
df <- clubdata_rsiena_freq[[4]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# classify dyads
dyadcount <- igraph::dyad.census(G1)
# add the total number of dyads to the graph
dyadcount$total <- (vcount(G1)*(vcount(G1)-1))/2
dyadcount#> $mut
#> [1] 7
#>
#> $asym
#> [1] 2
#>
#> $null
#> [1] 27
#>
#> $total
#> [1] 36# compare values with a random graph of the same size with the same density
dens <- igraph::graph.density(G1)
size <- igraph::vcount(G1)
trial <- 1000
recip <- rep(NA, trial)
for ( i in 1:trial ){
random_graph <- igraph::erdos.renyi.game(n = size, p.or.m = dens, directed = TRUE)
recip[i] <- igraph::dyad.census(random_graph)$mut
}
{hist(recip, main="number of reciprocated Kudos in random graph", xlab="", )
abline(v=dyadcount$mut, col="red", lwd=3)}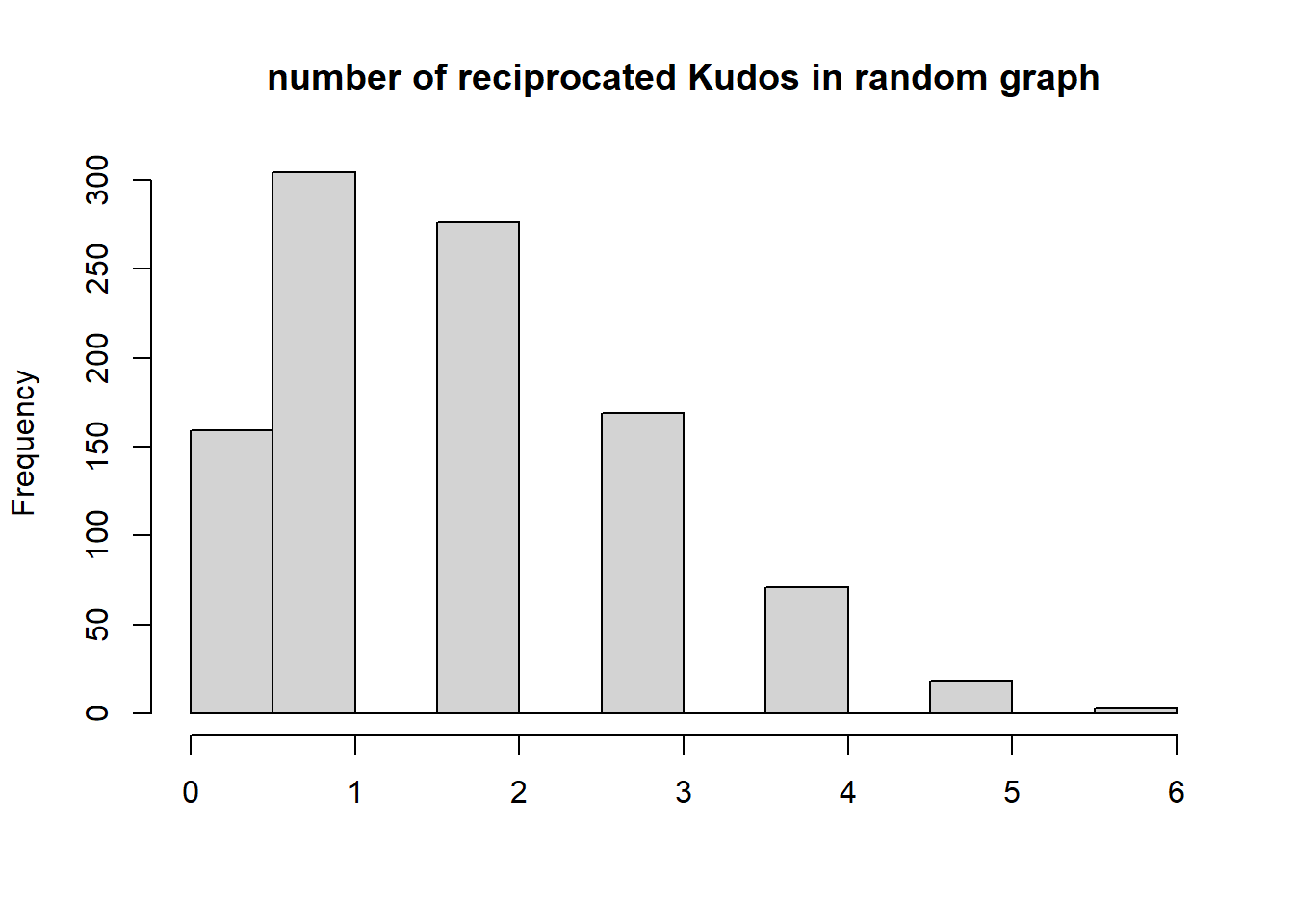
Club 5
# make igraph object for the club, at wave 1
df <- clubdata_rsiena_freq[[5]] # grab club
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# make an 'igraph object'
G1 <- igraph::graph_from_adjacency_matrix(knet1, mode = "directed", weighted = NULL, diag = TRUE, add.colnames = NA, add.rownames = NA)
# classify dyads
dyadcount <- igraph::dyad.census(G1)
# add the total number of dyads to the graph
dyadcount$total <- (vcount(G1)*(vcount(G1)-1))/2
dyadcount#> $mut
#> [1] 195
#>
#> $asym
#> [1] 119
#>
#> $null
#> [1] 2536
#>
#> $total
#> [1] 2850# compare values with a random graph of the same size with the same density
dens <- igraph::graph.density(G1)
size <- igraph::vcount(G1)
trial <- 1000
recip <- rep(NA, trial)
for ( i in 1:trial ){
random_graph <- igraph::erdos.renyi.game(n = size, p.or.m = dens, directed = TRUE)
recip[i] <- igraph::dyad.census(random_graph)$mut
}
{hist(recip, main="number of reciprocated Kudos in random graph", xlab="", )
abline(v=dyadcount$mut, col="red", lwd=3)}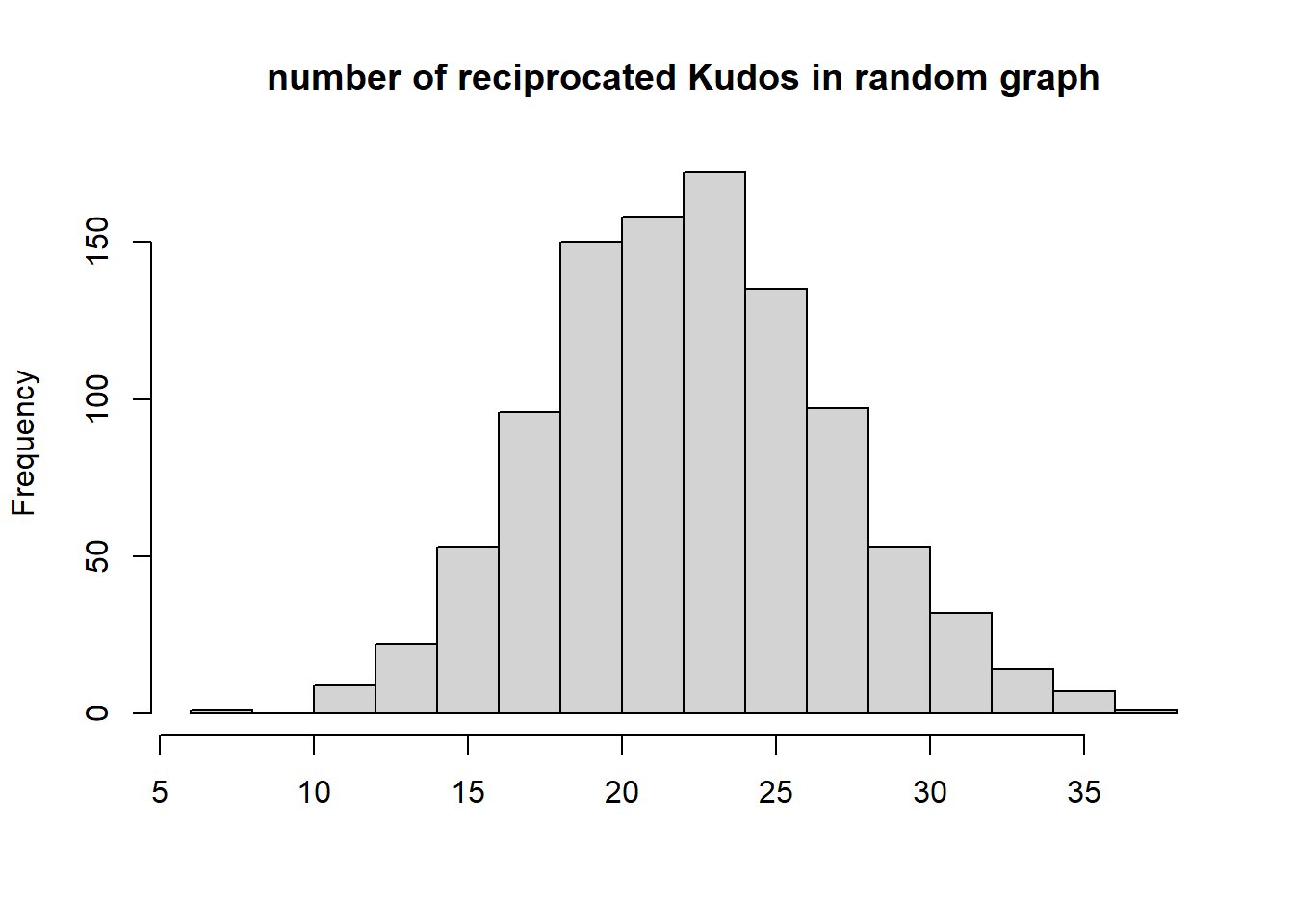
Conclusion: Kudos tend to be reciprocated!
Gender composition
Let’s investigate the gender composition of the club. We must retrieve gender from the object (note that we use the clubdata object, not the RSiena object). Then we make a categorical gender variable and plot it.
Club 1
df <- clubdata[[1]] # grab club
# retrieve node-attribute gender from object
male <- df$male
female <- df$female
other <- df$other
# as factor
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# make dataframe
df <- data.frame(
gender = as.factor(c("Male", "Female", "Other", "Missing")),
n = c(length(gender[gender == "Male"]), length(gender[gender == "Female"]), length(gender[gender == "Other"]), length(gender[gender == "Missing"])),
freq = c(round((length(gender[gender=="Male"])/length(gender) *100), digits=1), round((length(gender[gender=="Female"])/length(gender) *100), digits=1), round((length(gender[gender=="Other"])/length(gender)*100), digits=1), round((length(gender[gender=="Missing"])/length(gender)*100), digits=1))
)
# plot
df %>%
mutate(gender = fct_reorder(gender, -n)) %>%
ggplot(aes(gender, n, fill=gender)) +
geom_bar(stat="identity", width=0.8) +
geom_text(aes(label=paste0(freq,"%")), vjust=1.5, colour="white")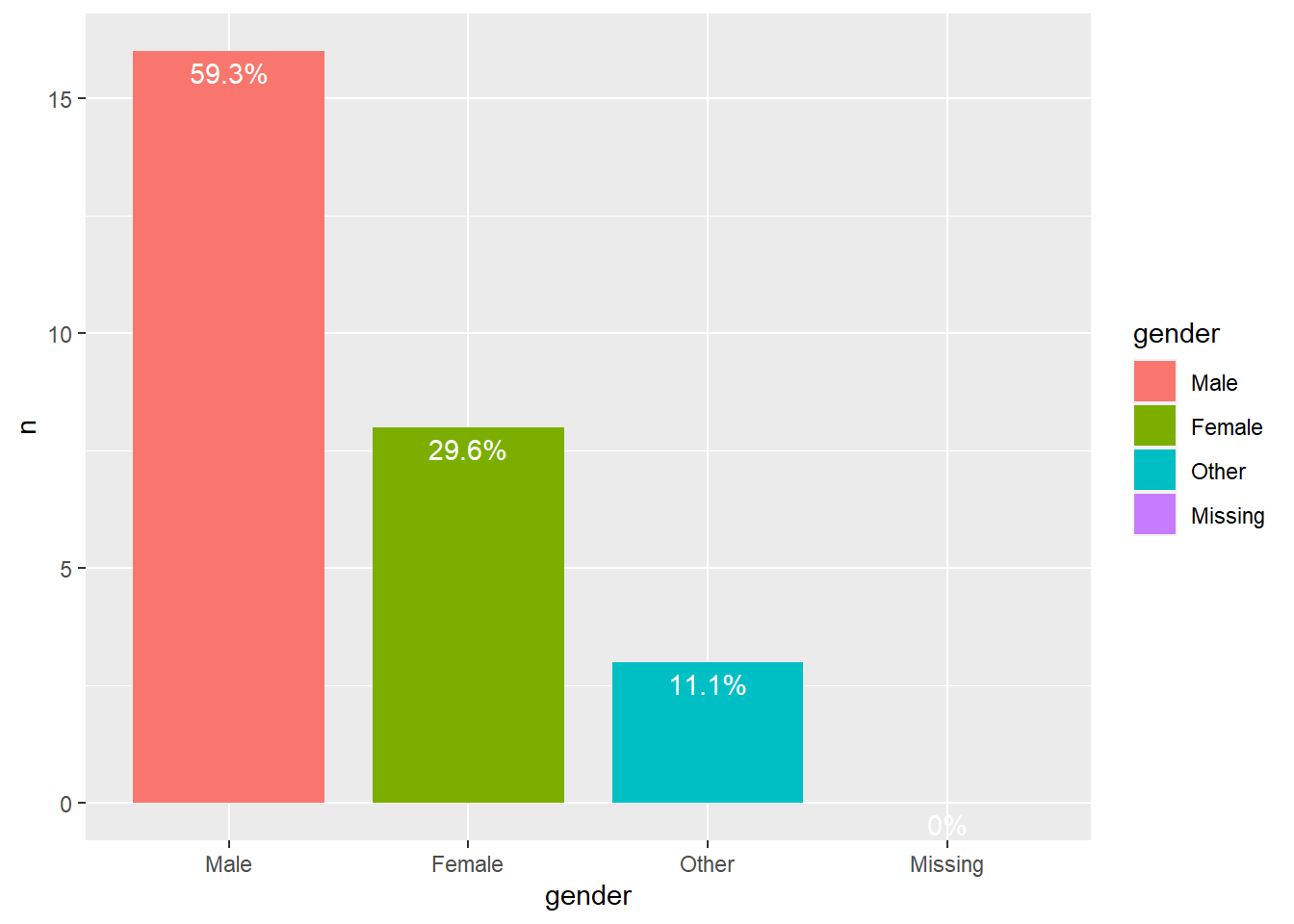
Club 2
df <- clubdata[[2]] # grab club
# retrieve node-attribute gender from object
male <- df$male
female <- df$female
other <- df$other
# as factor
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# make dataframe
df <- data.frame(
gender = as.factor(c("Male", "Female", "Other", "Missing")),
n = c(length(gender[gender == "Male"]), length(gender[gender == "Female"]), length(gender[gender == "Other"]), length(gender[gender == "Missing"])),
freq = c(round((length(gender[gender=="Male"])/length(gender) *100), digits=1), round((length(gender[gender=="Female"])/length(gender) *100), digits=1), round((length(gender[gender=="Other"])/length(gender)*100), digits=1), round((length(gender[gender=="Missing"])/length(gender)*100), digits=1))
)
# plot
df %>%
mutate(gender = fct_reorder(gender, -n)) %>%
ggplot(aes(gender, n, fill=gender)) +
geom_bar(stat="identity", width=0.8) +
geom_text(aes(label=paste0(freq,"%")), vjust=1.5, colour="white")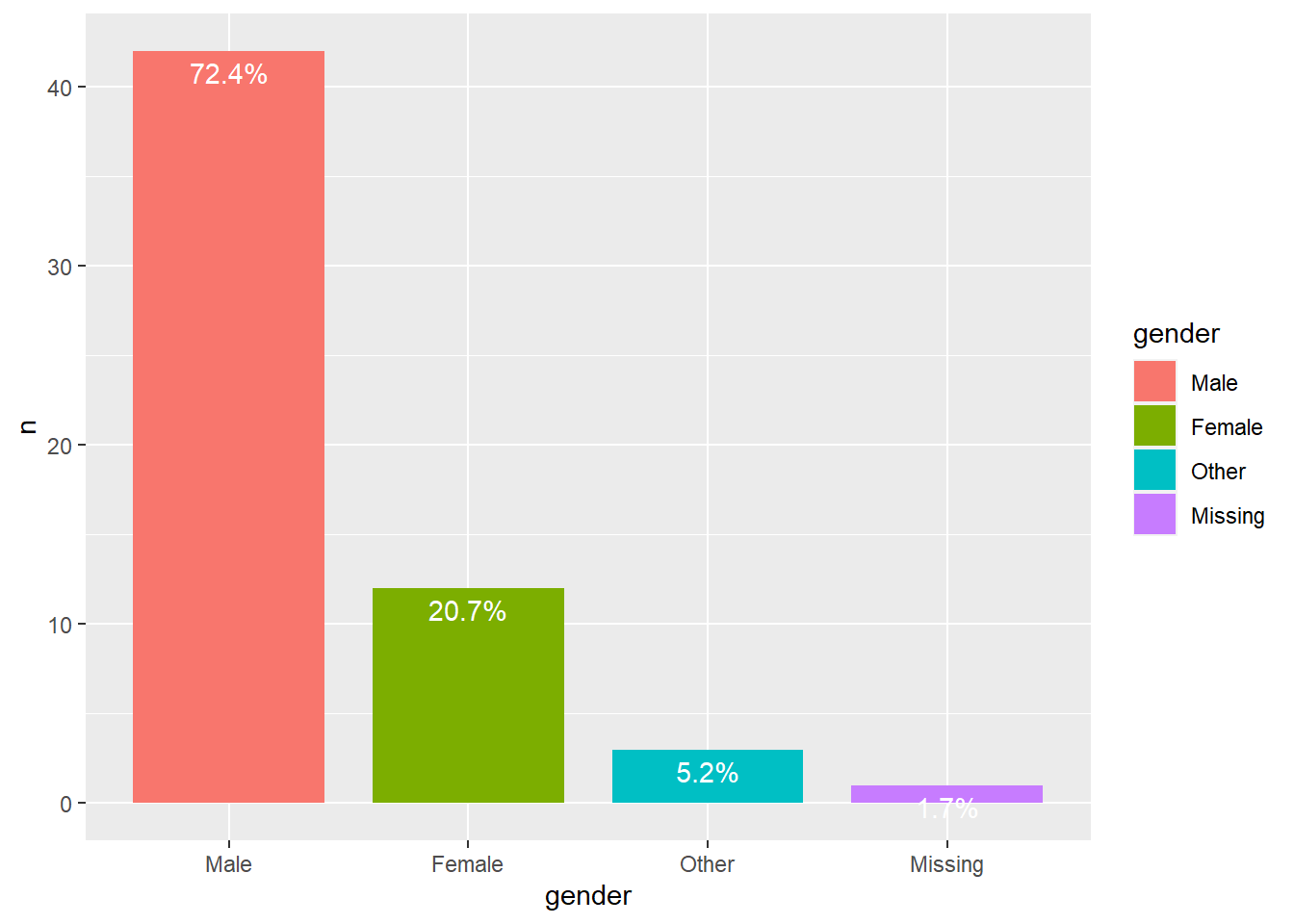
Club 3
df <- clubdata[[3]] # grab club
# retrieve node-attribute gender from object
male <- df$male
female <- df$female
other <- df$other
# as factor
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# make dataframe
df <- data.frame(
gender = as.factor(c("Male", "Female", "Other", "Missing")),
n = c(length(gender[gender == "Male"]), length(gender[gender == "Female"]), length(gender[gender == "Other"]), length(gender[gender == "Missing"])),
freq = c(round((length(gender[gender=="Male"])/length(gender) *100), digits=1), round((length(gender[gender=="Female"])/length(gender) *100), digits=1), round((length(gender[gender=="Other"])/length(gender)*100), digits=1), round((length(gender[gender=="Missing"])/length(gender)*100), digits=1))
)
# plot
df %>%
mutate(gender = fct_reorder(gender, -n)) %>%
ggplot(aes(gender, n, fill=gender)) +
geom_bar(stat="identity", width=0.8) +
geom_text(aes(label=paste0(freq,"%")), vjust=1.5, colour="white")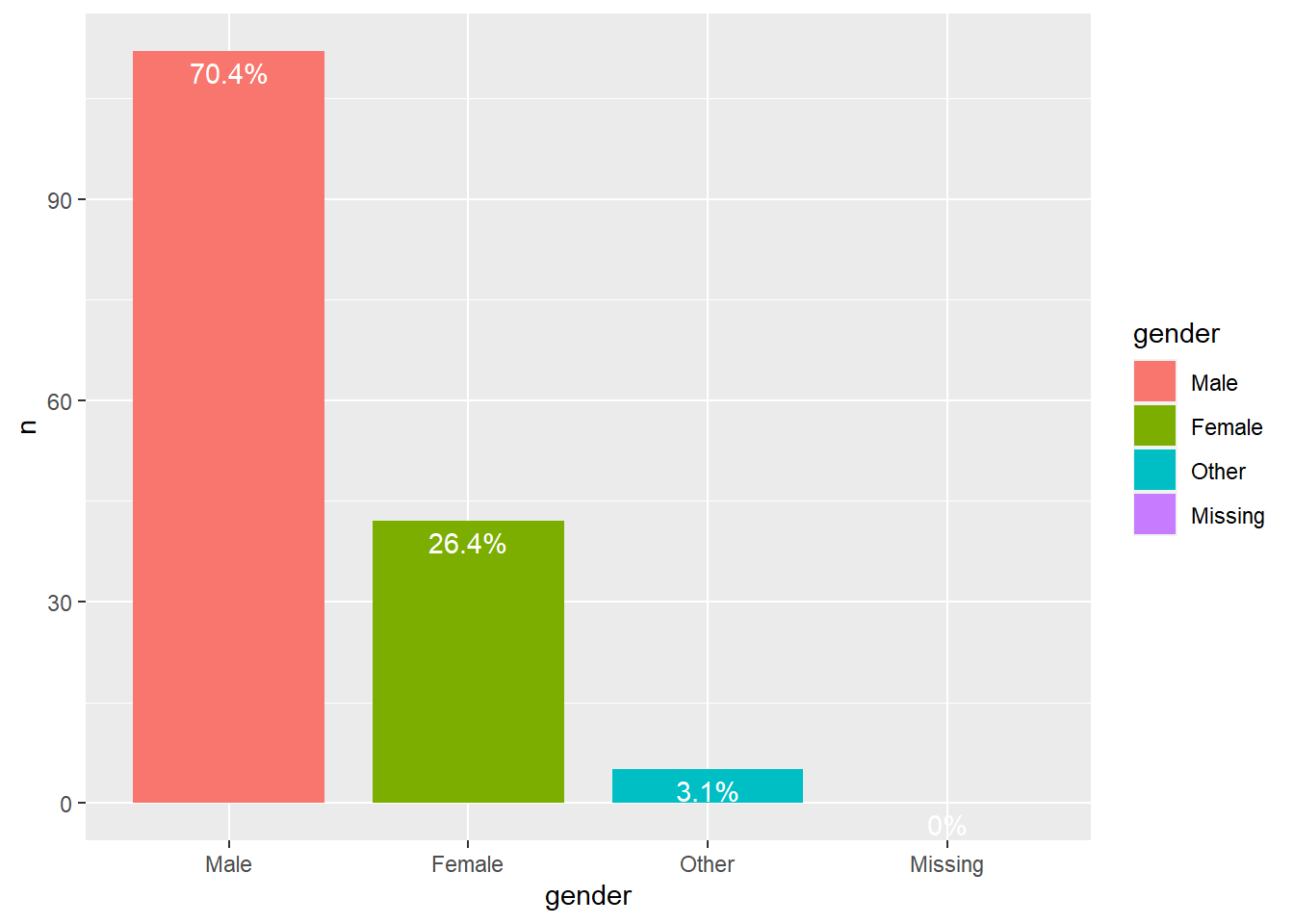
Club 4
df <- clubdata[[4]] # grab club
# retrieve node-attribute gender from object
male <- df$male
female <- df$female
other <- df$other
# as factor
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# make dataframe
df <- data.frame(
gender = as.factor(c("Male", "Female", "Other", "Missing")),
n = c(length(gender[gender == "Male"]), length(gender[gender == "Female"]), length(gender[gender == "Other"]), length(gender[gender == "Missing"])),
freq = c(round((length(gender[gender=="Male"])/length(gender) *100), digits=1), round((length(gender[gender=="Female"])/length(gender) *100), digits=1), round((length(gender[gender=="Other"])/length(gender)*100), digits=1), round((length(gender[gender=="Missing"])/length(gender)*100), digits=1))
)
# plot
df %>%
mutate(gender = fct_reorder(gender, -n)) %>%
ggplot(aes(gender, n, fill=gender)) +
geom_bar(stat="identity", width=0.8) +
geom_text(aes(label=paste0(freq,"%")), vjust=1.5, colour="white")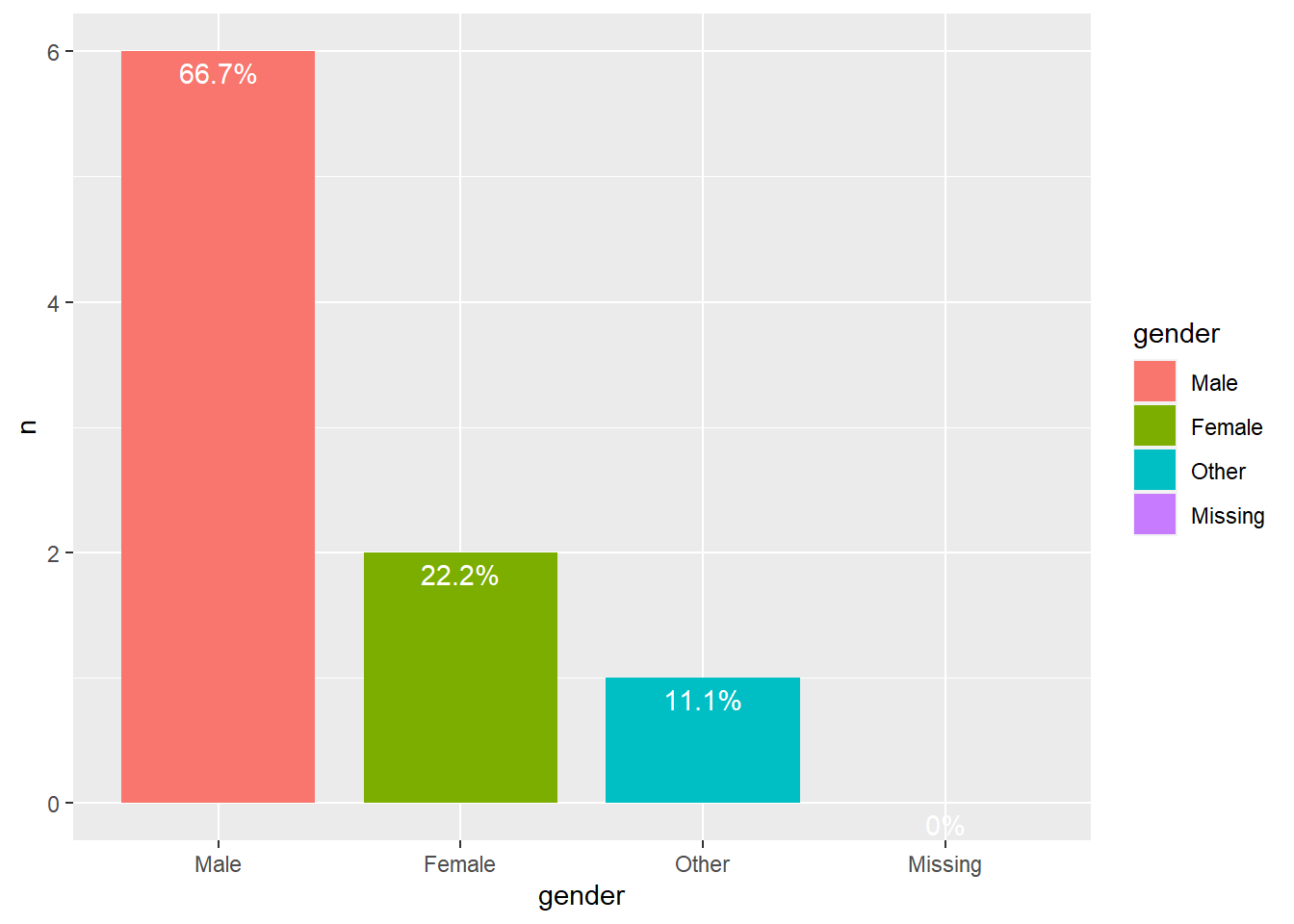
Club 5
df <- clubdata[[5]] # grab club
# retrieve node-attribute gender from object
male <- df$male
female <- df$female
other <- df$other
# as factor
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# make dataframe
df <- data.frame(
gender = as.factor(c("Male", "Female", "Other", "Missing")),
n = c(length(gender[gender == "Male"]), length(gender[gender == "Female"]), length(gender[gender == "Other"]), length(gender[gender == "Missing"])),
freq = c(round((length(gender[gender=="Male"])/length(gender) *100), digits=1), round((length(gender[gender=="Female"])/length(gender) *100), digits=1), round((length(gender[gender=="Other"])/length(gender)*100), digits=1), round((length(gender[gender=="Missing"])/length(gender)*100), digits=1))
)
# plot
df %>%
mutate(gender = fct_reorder(gender, -n)) %>%
ggplot(aes(gender, n, fill=gender)) +
geom_bar(stat="identity", width=0.8) +
geom_text(aes(label=paste0(freq,"%")), vjust=1.5, colour="white")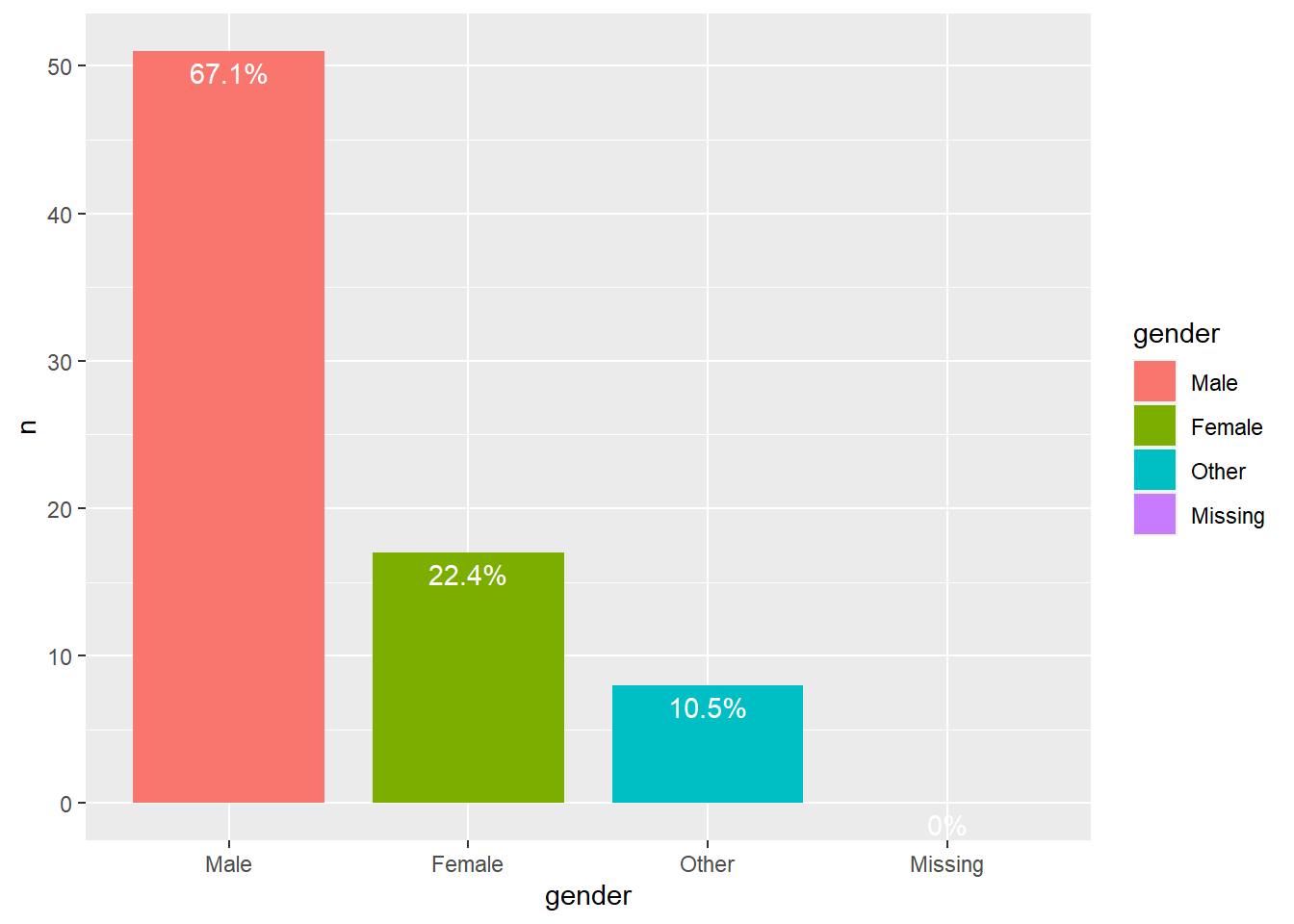
We can see that in all clubs men are the majority.
Gender segregation
Let’s now investigate segregation along gender in the kudos network.
Let’s start with describing the total density and intra- (same gender) and intergroup (different gender) densities. We also calculate the Coleman Homophily index for gender, which reflects gender segregation while taking into account the relative group size of gender categories.
Club 1
df <- clubdata_rsiena_freq[[1]] # grab club
df2 <- clubdata[[1]]
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# for some reason constructing the dyad-similarity matrix for gender with the rsiena object did not work, so we use the clubdata.RData.
male <- df2$male
female <- df2$female
other <- df2$other
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# construct dyad similarity matrix
gender_m <- fhomomat(gender)
# make object to store results
desmat <- matrix(NA, nrow=4, ncol=1)
# use functions
desmat[1, 1] <- fdensity(knet1)
desmat[2, 1] <- fdensityintra(knet1, gender_m)
desmat[3, 1] <- fdensityinter(knet1, gender_m)
desmat[4, 1] <- fscolnet(knet1, gender)
colnames(desmat) <- c("Kudos network")
rownames(desmat) <- c("total density", "same gender density", "different gender density", "Coleman's homophily index")
knitr::kable(desmat, digits=3, "html", caption="Gender segregation in friendship and kudo network") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| Kudos network | |
|---|---|
| total density | 0.118 |
| same gender density | 0.151 |
| different gender density | 0.086 |
| Coleman’s homophily index | 0.232 |
Club 2
df <- clubdata_rsiena_freq[[2]] # grab club
df2 <- clubdata[[2]]
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# for some reason constructing the dyad-similarity matrix for gender with the rsiena object did not work, so we use the clubdata.RData.
male <- df2$male
female <- df2$female
other <- df2$other
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# construct dyad similarity matrix
gender_m <- fhomomat(gender)
# make object to store results
desmat <- matrix(NA, nrow=4, ncol=1)
# use functions
desmat[1, 1] <- fdensity(knet1)
desmat[2, 1] <- fdensityintra(knet1, gender_m)
desmat[3, 1] <- fdensityinter(knet1, gender_m)
desmat[4, 1] <- fscolnet(knet1, gender)
colnames(desmat) <- c("Kudos network")
rownames(desmat) <- c("total density", "same gender density", "different gender density", "Coleman's homophily index")
knitr::kable(desmat, digits=3, "html", caption="Gender segregation in friendship and kudo network") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| Kudos network | |
|---|---|
| total density | 0.189 |
| same gender density | 0.250 |
| different gender density | 0.103 |
| Coleman’s homophily index | 0.343 |
Club 3
df <- clubdata_rsiena_freq[[3]] # grab club
df2 <- clubdata[[3]]
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# for some reason constructing the dyad-similarity matrix for gender with the rsiena object did not work, so we use the clubdata.RData.
male <- df2$male
female <- df2$female
other <- df2$other
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# construct dyad similarity matrix
gender_m <- fhomomat(gender)
# make object to store results
desmat <- matrix(NA, nrow=4, ncol=1)
# use functions
desmat[1, 1] <- fdensity(knet1)
desmat[2, 1] <- fdensityintra(knet1, gender_m)
desmat[3, 1] <- fdensityinter(knet1, gender_m)
desmat[4, 1] <- fscolnet(knet1, gender)
colnames(desmat) <- c("Kudos network")
rownames(desmat) <- c("total density", "same gender density", "different gender density", "Coleman's homophily index")
knitr::kable(desmat, digits=3, "html", caption="Gender segregation in friendship and kudo network") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| Kudos network | |
|---|---|
| total density | 0.017 |
| same gender density | 0.017 |
| different gender density | 0.018 |
| Coleman’s homophily index | 0.040 |
Club 4
df <- clubdata_rsiena_freq[[4]] # grab club
df2 <- clubdata[[4]]
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# for some reason constructing the dyad-similarity matrix for gender with the rsiena object did not work, so we use the clubdata.RData.
male <- df2$male
female <- df2$female
other <- df2$other
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# construct dyad similarity matrix
gender_m <- fhomomat(gender)
# make object to store results
desmat <- matrix(NA, nrow=4, ncol=1)
# use functions
desmat[1, 1] <- fdensity(knet1)
desmat[2, 1] <- fdensityintra(knet1, gender_m)
desmat[3, 1] <- fdensityinter(knet1, gender_m)
desmat[4, 1] <- fscolnet(knet1, gender)
colnames(desmat) <- c("Kudos network")
rownames(desmat) <- c("total density", "same gender density", "different gender density", "Coleman's homophily index")
knitr::kable(desmat, digits=3, "html", caption="Gender segregation in friendship and kudo network") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| Kudos network | |
|---|---|
| total density | 0.381 |
| same gender density | 0.286 |
| different gender density | 0.429 |
| Coleman’s homophily index | -0.347 |
Club 5
df <- clubdata_rsiena_freq[[5]] # grab club
df2 <- clubdata[[5]]
knet <- df$depvars$kudonet # take Kudo network
knet1 <- knet[,,1] # take wave 1 only for now
# for some reason constructing the dyad-similarity matrix for gender with the rsiena object did not work, so we use the clubdata.RData.
male <- df2$male
female <- df2$female
other <- df2$other
gender <- NA
gender <- ifelse(male == 1, "Male", gender)
gender <- ifelse(female == 1, "Female", gender)
gender <- ifelse(other == 1, "Other", gender)
gender <- ifelse(is.na(gender), "Missing", gender) # missing category
# construct dyad similarity matrix
gender_m <- fhomomat(gender)
# make object to store results
desmat <- matrix(NA, nrow=4, ncol=1)
# use functions
desmat[1, 1] <- fdensity(knet1)
desmat[2, 1] <- fdensityintra(knet1, gender_m)
desmat[3, 1] <- fdensityinter(knet1, gender_m)
desmat[4, 1] <- fscolnet(knet1, gender)
colnames(desmat) <- c("Kudos network")
rownames(desmat) <- c("total density", "same gender density", "different gender density", "Coleman's homophily index")
knitr::kable(desmat, digits=3, "html", caption="Gender segregation in friendship and kudo network") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| Kudos network | |
|---|---|
| total density | 0.122 |
| same gender density | 0.117 |
| different gender density | 0.129 |
| Coleman’s homophily index | 0.025 |
Density in the kudos network is not much higher within-gender than between-genders, though this ratio differs per club (and in one clubs between-gender kudos are more common).
Coleman’s Homophily indices vary across clubs. In some clubs there is slight gender segregation (or same-gender preference) in kudos ties (values higher than 0, but rather close to 0), though in others members tend to give kudos to others outside their own gender group (values lower than 0).
Behavior
We plotted the development of the mean of running attributes (Figure 3 of the manuscript). The script to replicate this plot can be found here.
Correlation between frequency and volume
Let’s plot the relation between frequency in times per week and volume in hours per week. We also calculate Kendall’s tau-b, i.e. a non-parametric measure of correlation on ranks (cf. Khamis (2008)).
Club 1
df <- clubdata[[1]] # grab club
df <- data.frame(x = as.matrix(df$time_run), y = as.matrix(df$freq_run))
c <- cor.test(df$x, df$y, method="kendall")
plot(df, type="b",
main = "Relation between weekly running variables: frequency and (half) hours",
sub = "Note: running frequency capped with a maximum of 7, hours with a maximum of 7",
xlab = "Hours per week", ylab = "sessions per week",
col="blue", lwd=c(1,rep(6,10000))) +
text(x = 1.7, y = 5, round(c$estimate, digits=2)) +
text(x = .7, y = 5, "Kendall's tau-b =")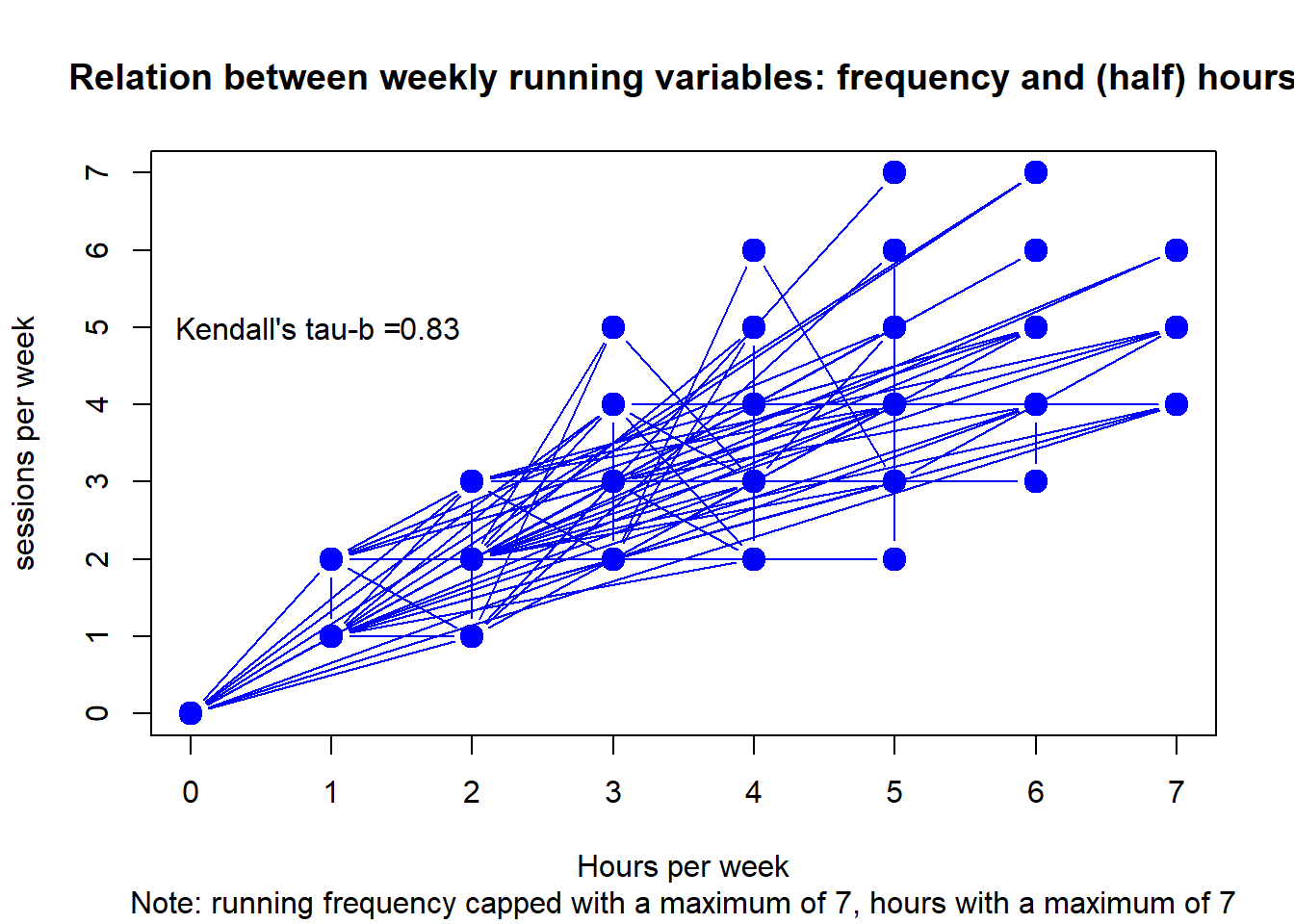
#> integer(0)Club 2
df <- clubdata[[2]] # grab club
df <- data.frame(x = as.matrix(df$time_run), y = as.matrix(df$freq_run))
c <- cor.test(df$x,df$y,method="kendall")
plot(df, type="b",
main = "Relation between weekly running variables: frequency and hours",
sub = "Note: running frequency capped with a maximum of 7, hours with a maximum of 7",
xlab = "hours per week", ylab = "sessions per week",
col="blue", lwd=c(1,rep(6,10000))) +
text(x = 1.7, y = 5, round(c$estimate, digits=2)) +
text(x = .7, y = 5, "Kendall's tau-b =")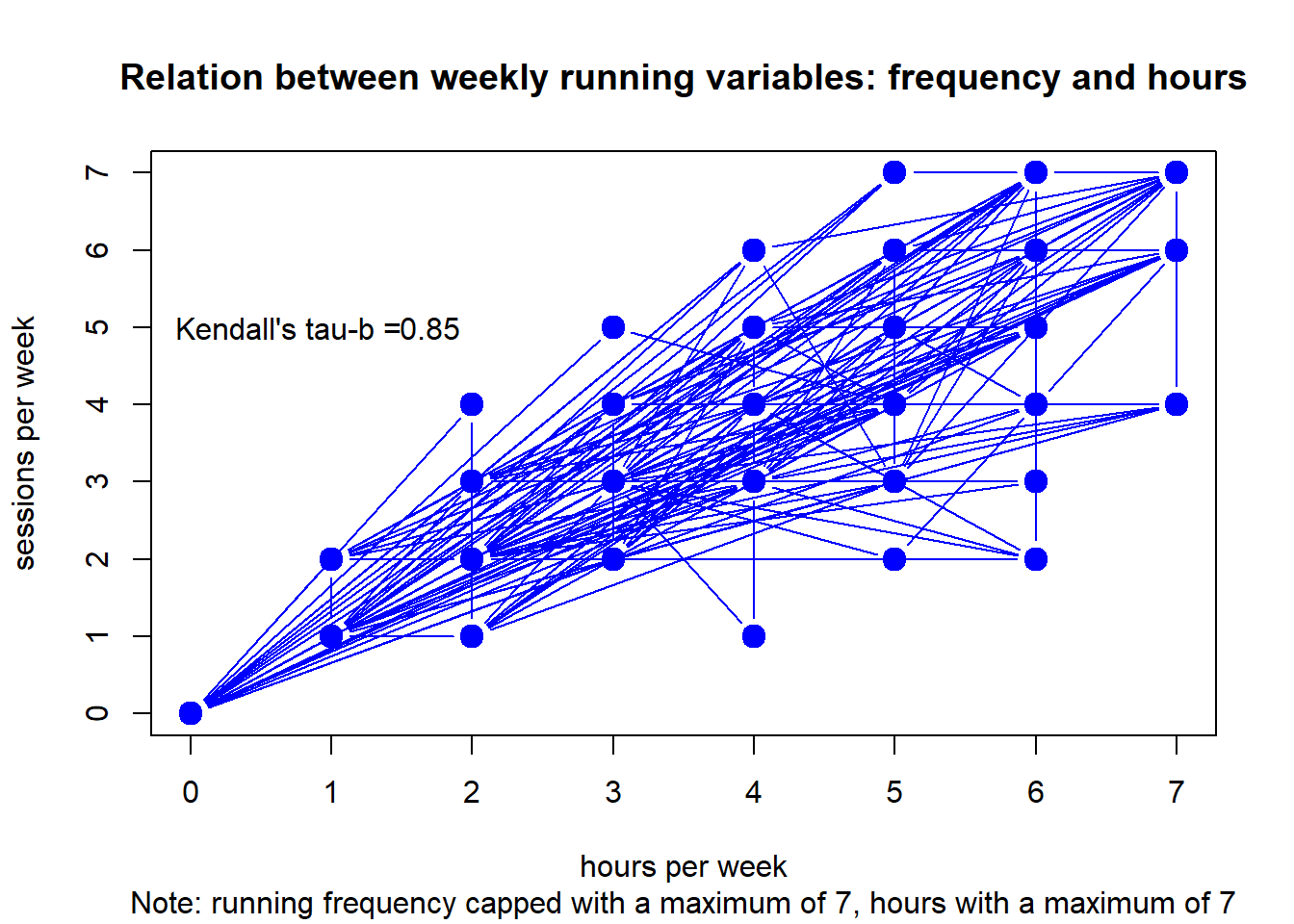
#> integer(0)Club 3
df <- clubdata[[3]] # grab club
df <- data.frame(x = as.matrix(df$time_run), y = as.matrix(df$freq_run))
c <- cor.test(df$x,df$y,method="kendall")
plot(df, type="b",
main = "Relation between weekly running variables: frequency and hours",
sub = "Note: running frequency capped with a maximum of 7, hours with a maximum of 7",
xlab = "hours per week", ylab = "sessions per week",
col="blue", lwd=c(1,rep(6,10000))) +
text(x = 1.7, y = 5, round(c$estimate, digits=2)) +
text(x = .7, y = 5, "Kendall's tau-b =")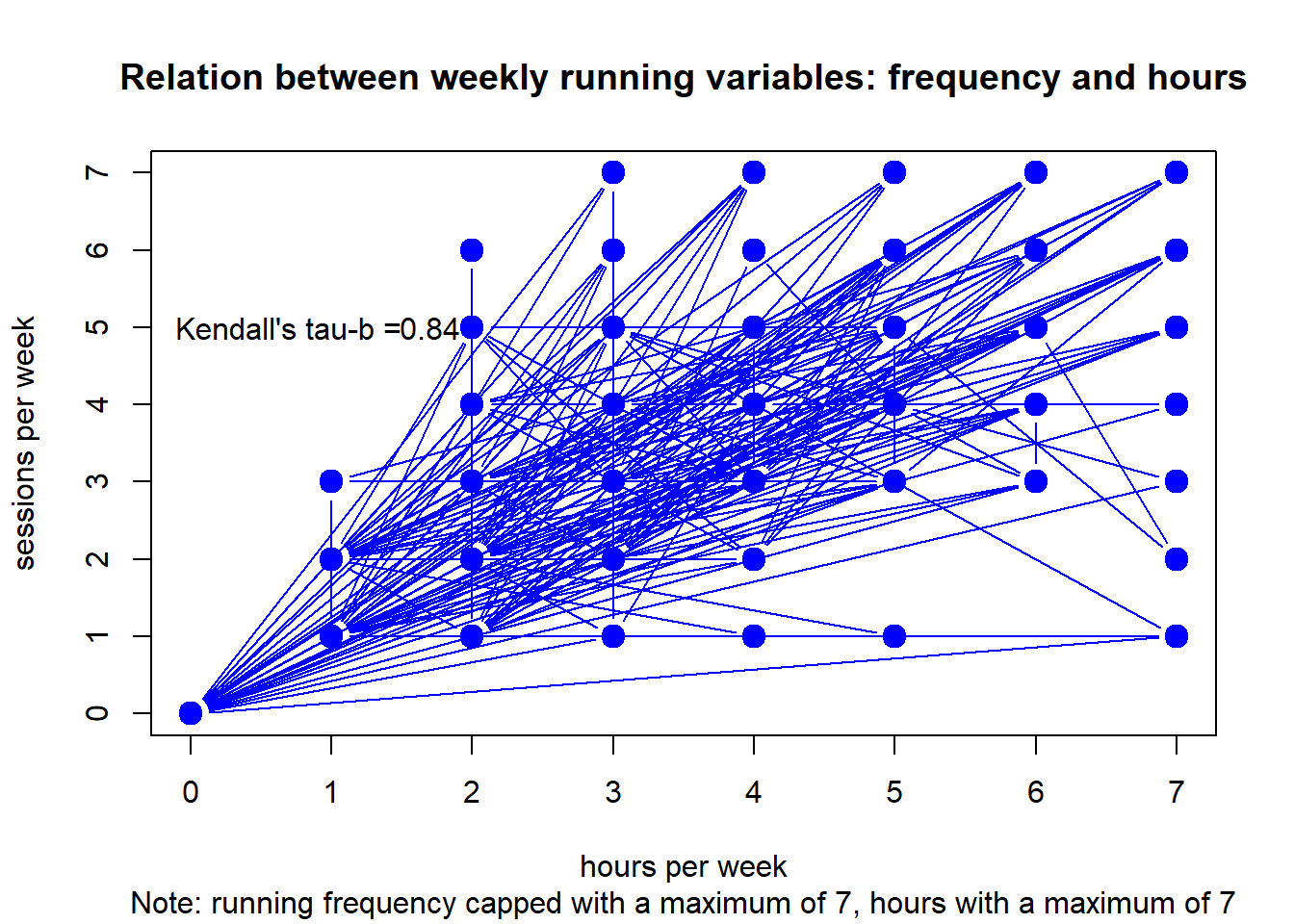
#> integer(0)Club 4
df <- clubdata[[4]] # grab club
df <- data.frame(x = as.matrix(df$time_run), y = as.matrix(df$freq_run))
c <- cor.test(df$x,df$y,method="kendall")
plot(df, type="b",
main = "Relation between weekly running variables: frequency and hours",
sub = "Note: running frequency capped with a maximum of 7, hours with a maximum of 7",
xlab = "hours per week", ylab = "sessions per week",
col="blue", lwd=c(1,rep(6,10000))) +
text(x = 1.5, y = 5, round(c$estimate, digits=2)) +
text(x = .7, y = 5, "Kendall's tau-b =")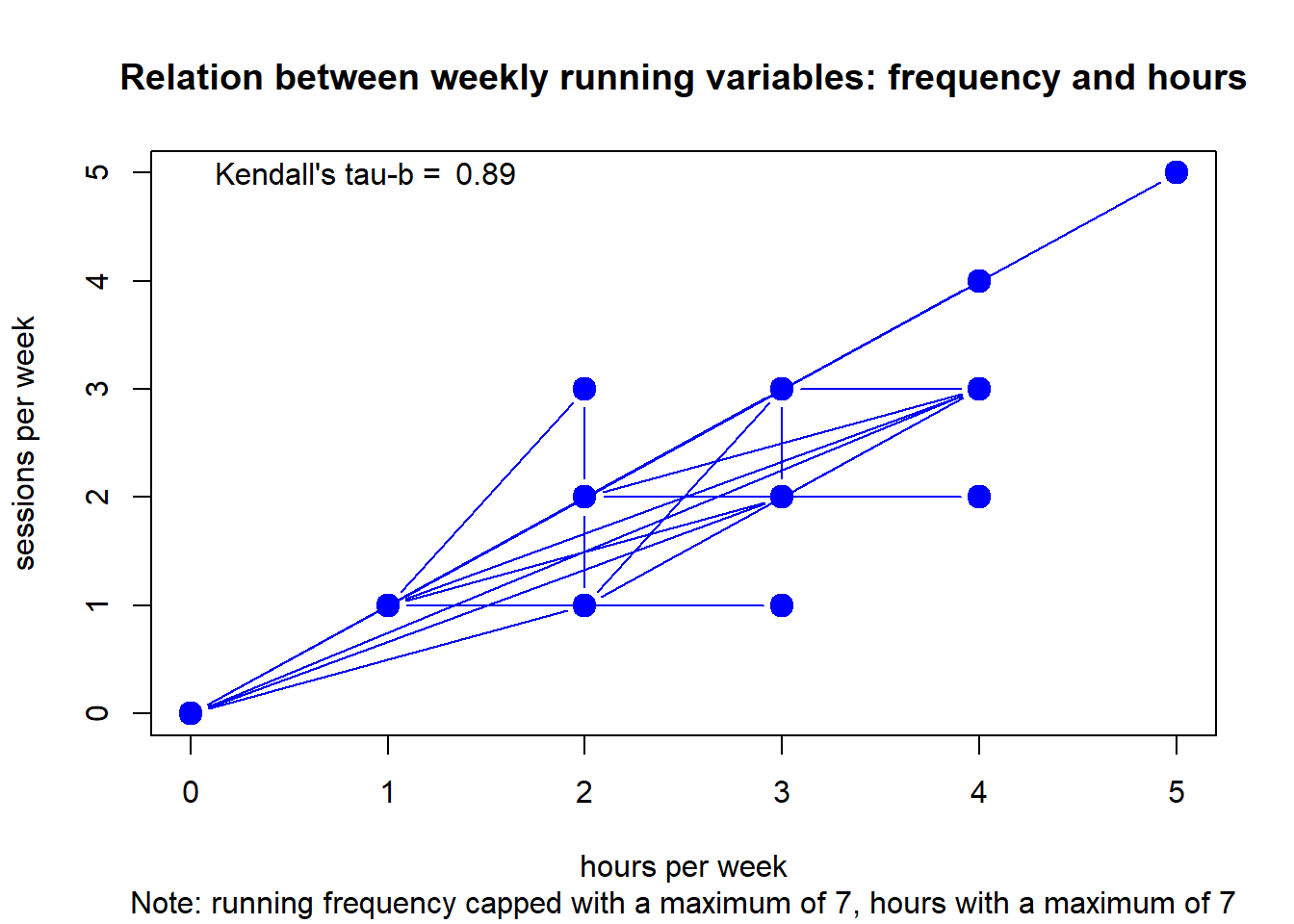
#> integer(0)Club 5
df <- clubdata[[5]] # grab club
df <- data.frame(x = as.matrix(df$time_run), y = as.matrix(df$freq_run))
c <- cor.test(df$x,df$y,method="kendall")
plot(df, type="b",
main = "Relation between weekly running variables: frequency and hours",
sub = "Note: running frequency capped with a maximum of 7, hours with a maximum of 7",
xlab = "hours per week", ylab = "sessions per week",
col="blue", lwd=c(1,rep(6,10000))) +
text(x = 2, y = 7, round(c$estimate, digits=2)) +
text(x = .7, y = 7, "Kendall's tau-b =")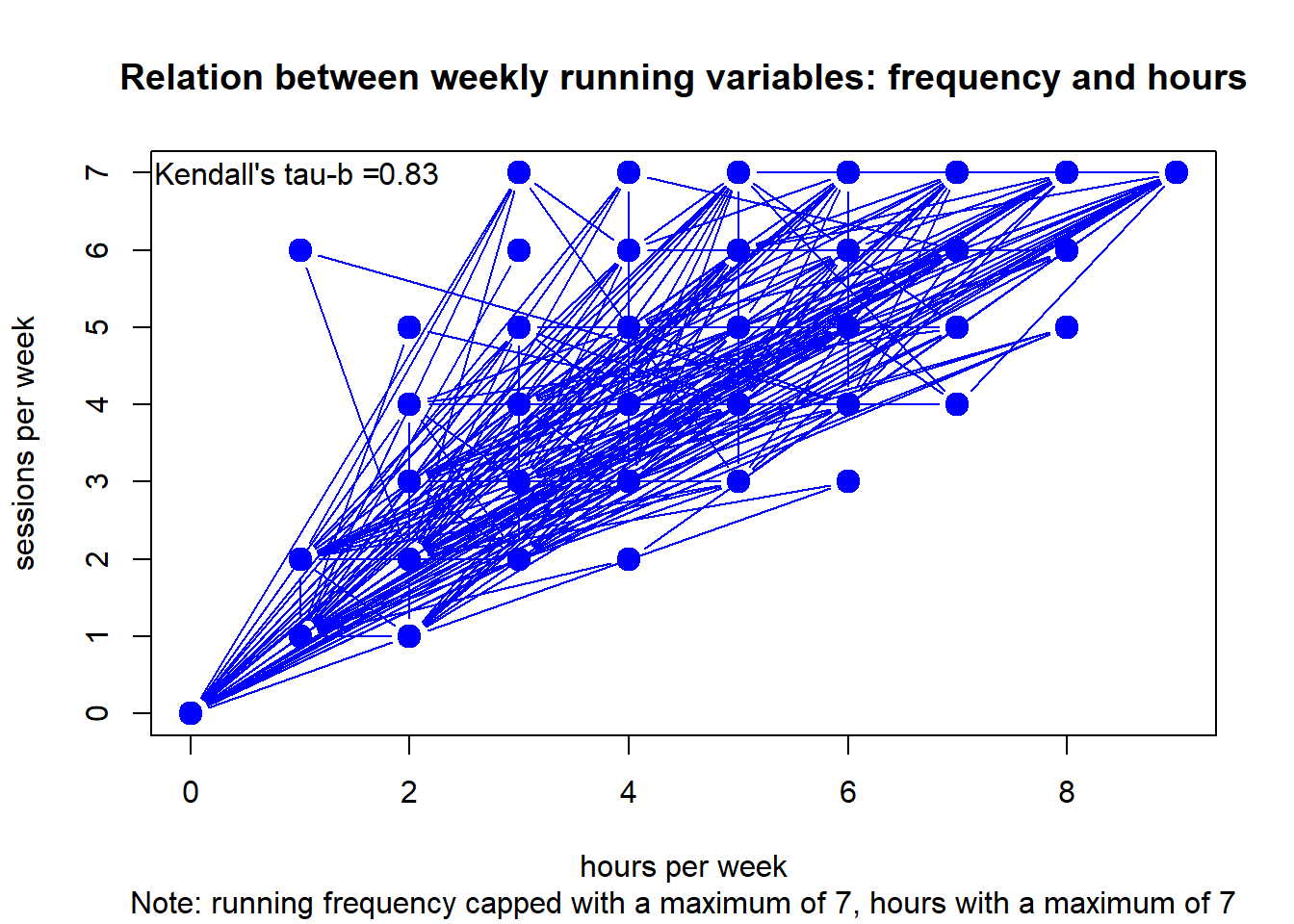
#> integer(0)Although frequency and duration are strongly related, there is some invariance.
Network autocorrelation
We have now covered the sport activity levels of our club-athletes, and the extent to which kudos-associations are segregated along gender. Last, we will explore if kudos-ties are also segregated along activity levels. Or in other words: do people with similar activity levels tend to socialize to a greater extent - by exchanging kudos - even when taking into account the opportunity structures for ‘interacting’ with (dis)similar others?
We use Moran’s I spatial autocorrelation measure for this, which is the correlation between the behavioral score of actor i and the (total/mean) behavioral score of alters j to whom i is connected directly. We also calculated Moran’s I by including the behavioral scores of the actors h to whom i is indirectly tied, and used the negative exponential function as described by Chen (2013) as a distance-decay function for assigning weights.
Club 1
df <- clubdata[[1]] # grab club
# get behavioral data
# at time t=1
t=1
freq1 <- df$freq_run[,,t] # running frequencies wave 1
vol1 <- df$time_run[,,t] # running volume wave 1
# exclude NAs
na <- which(is.na(freq1))
freq1 <- freq1[-na]
vol1 <- vol1[-na]
# get kudos net
knet1 <- df$kudo[,,t]
# exclude NAs
knet1 <- knet1[-na,-na]
# as network object
knet1 <- network::as.network(knet1)
# we include geodistances: shortest path lengths from i to j
geodistances <- sna::geodist(knet1, count.paths=T)
geodistances <- geodistances$gdist
# set the distance 'to yourself' to 'Inf'
diag(geodistances) <- Inf
# first calculate Moran's i for alters at distance 1.
weights1 <- geodistances == 1
# and use the negative exponential distance-decay function
weights2 <- exp(-geodistances)
# calculate Moran's I
# for distance-1 and with distance decay, in the kudos network, for frequency and volume respectively
# we do not row standardize!
Mfreq <- fMoran.I(freq1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mfreq_distd <- fMoran.I(freq1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
Mvol <- fMoran.I(vol1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mvol_distd <- fMoran.I(vol1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
# make object to store results
# 1. frequency
f_mat <- matrix(NA, nrow=2, ncol=4)
f_mat[1,1] <- Mfreq$observed
f_mat[1,2] <- Mfreq$expected
f_mat[1,3] <- Mfreq$sd
f_mat[1,4] <- Mfreq$p.value
f_mat[2,1] <- Mfreq_distd$observed
f_mat[2,2] <- Mfreq_distd$expected
f_mat[2,3] <- Mfreq_distd$sd
f_mat[2,4] <- Mfreq_distd$p.value
# 2. volume
v_mat <- matrix(NA, nrow=2, ncol=4)
v_mat[1,1] <- Mvol$observed
v_mat[1,2] <- Mvol$expected
v_mat[1,3] <- Mvol$sd
v_mat[1,4] <- Mvol$p.value
v_mat[2,1] <- Mvol_distd$observed
v_mat[2,2] <- Mvol_distd$expected
v_mat[2,3] <- Mvol_distd$sd
v_mat[2,4] <- Mvol_distd$p.value
colnames(f_mat) <- colnames(v_mat) <- c("observed", "expected", "sd", "p-value")
rownames(f_mat) <- rownames(v_mat) <- c("direct kudos ties", "direct and indirect kudos ties (distance-decay)")
knitr::kable(f_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and weekly running frequency") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.01 | -0.04 | 0.13 | 0.66 |
| direct and indirect kudos ties (distance-decay) | 0.00 | -0.04 | 0.08 | 0.57 |
knitr::kable(v_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and monthly running volume") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | -0.01 | -0.04 | 0.13 | 0.80 |
| direct and indirect kudos ties (distance-decay) | -0.01 | -0.04 | 0.08 | 0.71 |
Club 2
df <- clubdata[[2]] # grab club
# get behavioral data
# at time t=1
t=1
freq1 <- df$freq_run[,,t] # running frequencies wave 1
vol1 <- df$time_run[,,t] # running volume wave 1
# exclude NAs
na <- which(is.na(freq1))
freq1 <- freq1[-na]
vol1 <- vol1[-na]
# get kudos net
knet1 <- df$kudo[,,t]
# exclude NAs
knet1 <- knet1[-na,-na]
# as network object
knet1 <- network::as.network(knet1)
# we include geodistances: shortest path lengths from i to j
geodistances <- sna::geodist(knet1, count.paths=T)
geodistances <- geodistances$gdist
# set the distance 'to yourself' to 'Inf'
diag(geodistances) <- Inf
# first calculate Moran's i for alters at distance 1.
weights1 <- geodistances == 1
# and use the negative exponential distance-decay function
weights2 <- exp(-geodistances)
# calculate Moran's I
# for distance-1 and with distance decay, in the kudos network, for frequency and volume respectively
# we do not row standardize!
Mfreq <- fMoran.I(freq1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mfreq_distd <- fMoran.I(freq1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
Mvol <- fMoran.I(vol1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mvol_distd <- fMoran.I(vol1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
# make object to store results
# 1. frequency
f_mat <- matrix(NA, nrow=2, ncol=4)
f_mat[1,1] <- Mfreq$observed
f_mat[1,2] <- Mfreq$expected
f_mat[1,3] <- Mfreq$sd
f_mat[1,4] <- Mfreq$p.value
f_mat[2,1] <- Mfreq_distd$observed
f_mat[2,2] <- Mfreq_distd$expected
f_mat[2,3] <- Mfreq_distd$sd
f_mat[2,4] <- Mfreq_distd$p.value
# 2. volume
v_mat <- matrix(NA, nrow=2, ncol=4)
v_mat[1,1] <- Mvol$observed
v_mat[1,2] <- Mvol$expected
v_mat[1,3] <- Mvol$sd
v_mat[1,4] <- Mvol$p.value
v_mat[2,1] <- Mvol_distd$observed
v_mat[2,2] <- Mvol_distd$expected
v_mat[2,3] <- Mvol_distd$sd
v_mat[2,4] <- Mvol_distd$p.value
colnames(f_mat) <- colnames(v_mat) <- c("observed", "expected", "sd", "p-value")
rownames(f_mat) <- rownames(v_mat) <- c("direct kudos ties", "direct and indirect kudos ties (distance-decay)")
knitr::kable(f_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and weekly running frequency") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.17 | -0.02 | 0.04 | 0 |
| direct and indirect kudos ties (distance-decay) | 0.05 | -0.02 | 0.01 | 0 |
knitr::kable(v_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and monthly running volume") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.21 | -0.02 | 0.04 | 0 |
| direct and indirect kudos ties (distance-decay) | 0.06 | -0.02 | 0.01 | 0 |
Club 3
df <- clubdata[[3]] # grab club
# get behavioral data
# at time t=1
t=1
freq1 <- df$freq_run[,,t] # running frequencies wave 1
vol1 <- df$time_run[,,t] # running volume wave 1
# exclude NAs
na <- which(is.na(freq1))
freq1 <- freq1[-na]
vol1 <- vol1[-na]
# get kudos net
knet1 <- df$kudo[,,t]
# exclude NAs
knet1 <- knet1[-na,-na]
# as network object
knet1 <- network::as.network(knet1)
# we include geodistances: shortest path lengths from i to j
geodistances <- sna::geodist(knet1, count.paths=T)
geodistances <- geodistances$gdist
# set the distance 'to yourself' to 'Inf'
diag(geodistances) <- Inf
# first calculate Moran's i for alters at distance 1.
weights1 <- geodistances == 1
# and use the negative exponential distance-decay function
weights2 <- exp(-geodistances)
# calculate Moran's I
# for distance-1 and with distance decay, in the kudos network, for frequency and volume respectively
# we do not row standardize!
Mfreq <- fMoran.I(freq1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mfreq_distd <- fMoran.I(freq1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
Mvol <- fMoran.I(vol1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mvol_distd <- fMoran.I(vol1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
# make object to store results
# 1. frequency
f_mat <- matrix(NA, nrow=2, ncol=4)
f_mat[1,1] <- Mfreq$observed
f_mat[1,2] <- Mfreq$expected
f_mat[1,3] <- Mfreq$sd
f_mat[1,4] <- Mfreq$p.value
f_mat[2,1] <- Mfreq_distd$observed
f_mat[2,2] <- Mfreq_distd$expected
f_mat[2,3] <- Mfreq_distd$sd
f_mat[2,4] <- Mfreq_distd$p.value
# 2. volume
v_mat <- matrix(NA, nrow=2, ncol=4)
v_mat[1,1] <- Mvol$observed
v_mat[1,2] <- Mvol$expected
v_mat[1,3] <- Mvol$sd
v_mat[1,4] <- Mvol$p.value
v_mat[2,1] <- Mvol_distd$observed
v_mat[2,2] <- Mvol_distd$expected
v_mat[2,3] <- Mvol_distd$sd
v_mat[2,4] <- Mvol_distd$p.value
colnames(f_mat) <- colnames(v_mat) <- c("observed", "expected", "sd", "p-value")
rownames(f_mat) <- rownames(v_mat) <- c("direct kudos ties", "direct and indirect kudos ties (distance-decay)")
knitr::kable(f_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and weekly running frequency") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.03 | -0.01 | 0.07 | 0.63 |
| direct and indirect kudos ties (distance-decay) | 0.02 | -0.01 | 0.03 | 0.28 |
knitr::kable(v_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and monthly running volume") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.16 | -0.01 | 0.07 | 0.02 |
| direct and indirect kudos ties (distance-decay) | 0.07 | -0.01 | 0.03 | 0.01 |
Club 4
df <- clubdata[[4]] # grab club
# get behavioral data
# at time t=1
t=1
freq1 <- df$freq_run[,,t] # running frequencies wave 1
vol1 <- df$time_run[,,t] # running volume wave 1
# exclude NAs
na <- which(is.na(freq1))
freq1 <- freq1[-na]
vol1 <- vol1[-na]
# get kudos net
knet1 <- df$kudo[,,t]
# exclude NAs
knet1 <- knet1[-na,-na]
# as network object
knet1 <- network::as.network(knet1)
# we include geodistances: shortest path lengths from i to j
geodistances <- sna::geodist(knet1, count.paths=T)
geodistances <- geodistances$gdist
# set the distance 'to yourself' to 'Inf'
diag(geodistances) <- Inf
# first calculate Moran's i for alters at distance 1.
weights1 <- geodistances == 1
# and use the negative exponential distance-decay function
weights2 <- exp(-geodistances)
# calculate Moran's I
# for distance-1 and with distance decay, in the kudos network, for frequency and volume respectively
# we do not row standardize!
Mfreq <- fMoran.I(freq1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mfreq_distd <- fMoran.I(freq1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
Mvol <- fMoran.I(vol1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mvol_distd <- fMoran.I(vol1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
# make object to store results
# 1. frequency
f_mat <- matrix(NA, nrow=2, ncol=4)
f_mat[1,1] <- Mfreq$observed
f_mat[1,2] <- Mfreq$expected
f_mat[1,3] <- Mfreq$sd
f_mat[1,4] <- Mfreq$p.value
f_mat[2,1] <- Mfreq_distd$observed
f_mat[2,2] <- Mfreq_distd$expected
f_mat[2,3] <- Mfreq_distd$sd
f_mat[2,4] <- Mfreq_distd$p.value
# 2. volume
v_mat <- matrix(NA, nrow=2, ncol=4)
v_mat[1,1] <- Mvol$observed
v_mat[1,2] <- Mvol$expected
v_mat[1,3] <- Mvol$sd
v_mat[1,4] <- Mvol$p.value
v_mat[2,1] <- Mvol_distd$observed
v_mat[2,2] <- Mvol_distd$expected
v_mat[2,3] <- Mvol_distd$sd
v_mat[2,4] <- Mvol_distd$p.value
colnames(f_mat) <- colnames(v_mat) <- c("observed", "expected", "sd", "p-value")
rownames(f_mat) <- rownames(v_mat) <- c("direct kudos ties", "direct and indirect kudos ties (distance-decay)")
knitr::kable(f_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and weekly running frequency") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | -0.31 | -0.17 | 0.26 | 0.57 |
| direct and indirect kudos ties (distance-decay) | -0.34 | -0.17 | 0.25 | 0.49 |
knitr::kable(v_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and monthly running volume") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | -0.28 | -0.17 | 0.28 | 0.69 |
| direct and indirect kudos ties (distance-decay) | -0.30 | -0.17 | 0.28 | 0.63 |
Club 5
df <- clubdata[[5]] # grab club
# get behavioral data
# at time t=1
t=1
freq1 <- df$freq_run[,,t] # running frequencies wave 1
vol1 <- df$time_run[,,t] # running volume wave 1
# exclude NAs
na <- which(is.na(freq1))
freq1 <- freq1[-na]
vol1 <- vol1[-na]
# get kudos net
knet1 <- df$kudo[,,t]
# exclude NAs
knet1 <- knet1[-na,-na]
# as network object
knet1 <- network::as.network(knet1)
# we include geodistances: shortest path lengths from i to j
geodistances <- sna::geodist(knet1, count.paths=T)
geodistances <- geodistances$gdist
# set the distance 'to yourself' to 'Inf'
diag(geodistances) <- Inf
# first calculate Moran's i for alters at distance 1.
weights1 <- geodistances == 1
# and use the negative exponential distance-decay function
weights2 <- exp(-geodistances)
# calculate Moran's I
# for distance-1 and with distance decay, in the kudos network, for frequency and volume respectively
# we do not row standardize!
Mfreq <- fMoran.I(freq1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mfreq_distd <- fMoran.I(freq1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
Mvol <- fMoran.I(vol1, scaled = FALSE, weight = weights1, na.rm = TRUE, rowstandardize = FALSE)
Mvol_distd <- fMoran.I(vol1, scaled = FALSE, weight = weights2, na.rm = TRUE, rowstandardize = FALSE)
# make object to store results
# 1. frequency
f_mat <- matrix(NA, nrow=2, ncol=4)
f_mat[1,1] <- Mfreq$observed
f_mat[1,2] <- Mfreq$expected
f_mat[1,3] <- Mfreq$sd
f_mat[1,4] <- Mfreq$p.value
f_mat[2,1] <- Mfreq_distd$observed
f_mat[2,2] <- Mfreq_distd$expected
f_mat[2,3] <- Mfreq_distd$sd
f_mat[2,4] <- Mfreq_distd$p.value
# 2. volume
v_mat <- matrix(NA, nrow=2, ncol=4)
v_mat[1,1] <- Mvol$observed
v_mat[1,2] <- Mvol$expected
v_mat[1,3] <- Mvol$sd
v_mat[1,4] <- Mvol$p.value
v_mat[2,1] <- Mvol_distd$observed
v_mat[2,2] <- Mvol_distd$expected
v_mat[2,3] <- Mvol_distd$sd
v_mat[2,4] <- Mvol_distd$p.value
colnames(f_mat) <- colnames(v_mat) <- c("observed", "expected", "sd", "p-value")
rownames(f_mat) <- rownames(v_mat) <- c("direct kudos ties", "direct and indirect kudos ties (distance-decay)")
knitr::kable(f_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and weekly running frequency") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.21 | -0.02 | 0.05 | 0 |
| direct and indirect kudos ties (distance-decay) | 0.09 | -0.02 | 0.02 | 0 |
knitr::kable(v_mat, digits=2, "html", caption="Moran's I statistic for spatial autocorrelation based on geodistances and monthly running volume") %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover"))| observed | expected | sd | p-value | |
|---|---|---|---|---|
| direct kudos ties | 0.25 | -0.02 | 0.05 | 0 |
| direct and indirect kudos ties (distance-decay) | 0.10 | -0.02 | 0.02 | 0 |
Here, the Moran’s I statistic tests whether club members that are closer to one another (i.e., having a shorter geodesic/path length), are more a similar with respect to their behavior, under the null hypothesis that behavior is ‘randomly distributed’ among the club members.
We observe that, indeed, kudos-ties that are closer to one another are more alike, with respect to running. Autocorrelation was stronger without the distance-decay function, which suggests that especially close alters (with path length one) are similar.
References
Copyright © 2021 Rob Franken